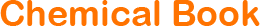
"SK-OV-3人卵巢癌细胞全年复苏|已有STR图谱
传代比例:1:2-1:4(首次传代建议1:2)
生长特性:贴壁生长
【细胞培养经验分享】启蒙老师的重要性:一般进实验室都有师兄师姐带着做,他们就是你做细胞的启蒙老师。他们的操作手法、细节、理论讲解就成了你操作的准则,如营养液、细胞瓶的摆放位置、灭菌处理程序、开盖手法、细胞吹打手法等等。要学会他们的正确操作,在第一次的时候就要重视。像养孩子一样养细胞,细胞有时真的很脆弱,最好每天都去看看它,以防止出现培养箱缺水、缺二氧化碳、停电、温度不够等异常现象,也好及时解决这些意外,避免重复实验带来的更大痛苦。好细胞要及时保种:细胞要分批传代,这样即使有一批出了问题,还有一批备用的。像后者一般人可能不容易做到。但这是我血的教训,有一次细胞污染了,全军覆没。当时可后悔没有保种。细胞跟人一样,不同的细胞,培养特性是不一样的。培养过程中要细细体会,不同细胞系使用不同的培养基和血清。
换液周期:每周2-3次
G361mel Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代,2-3天换液1次。;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:VP 267细胞、HCC-202细胞、HPB/ALL细胞
SK-OV-433 Cells;背景说明:卵巢癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:KYSE-450细胞、WIL2S细胞、DMS 273细胞
NCI-H2126 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代,每周2-3次。;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:NCIH1651细胞、BNL-HCC细胞、H-441细胞
背景信息:SK-OV-3由G.Trempe和L.J.Old在1973年从卵巢肿瘤病人的腹水分离得到。 此细胞对肿瘤坏死因子和几种细胞毒性药物包括白喉毒素、顺铂和阿霉素均耐受。 在裸鼠中致瘤,且形成与卵巢原位癌一致的中度分化的腺癌。
SK-OV-3人卵巢癌细胞全年复苏|已有STR图谱
产品包装:复苏发货:T25培养瓶(一瓶)或冻存发货:1ml冻存管(两支)
DSMZ菌株保藏中心成立于1969年,是德国的国家菌种保藏中心。该中心一直致力于细菌、真菌、质粒、抗菌素、人体和动物细胞、植物病毒等的分类、鉴定和保藏工作。DSMZ菌种保藏中心是欧洲规模最大的生物资源中心,保藏有动物细胞500多株。Riken BRC成立于1920年,是英国的国家菌种保藏中心。该中心一直致力于细菌、真菌、植物病毒等的分类、鉴定和保藏工作。日本Riken BRC(Riken生物资源保藏中心)是全球三大典型培养物收集中心之一。Riken保藏中心提供了很多细胞系。在世界范围内,这些细胞系,都在医学、科学和兽医中具有重要意义。Riken生物资源中心支持了各种学术、健康、食品和兽医机构的研究工作,并在世界各地不同组织的微生物实验室和研究机构中使用。
CO115 Cells;背景说明:结肠腺癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:SK.MEL.5细胞、PECAPJ34细胞、MADB-106细胞
IPEC-1 Cells;背景说明:小肠;上皮细胞;自发永生;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:P3 (Jiyoye)细胞、CCRF/CEM/0细胞、SKNO1细胞
J-111 Cells;背景说明:单核细胞白血病;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:Vero 76 clone E6细胞、LAD2细胞、HT-22细胞
225.28S Cells(提供STR鉴定图谱)
来源说明:细胞主要来源ATCC、ECACC、DSMZ、RIKEN等细胞库
物种来源:人源、鼠源等其它物种来源
SK-OV-3人卵巢癌细胞全年复苏|已有STR图谱
形态特性:上皮细胞样
细胞传代培养实验:体外培养的原代细胞或细胞株要在体外持续地培养就必须传代,以便获得稳定的细胞株或得到大量的同种细胞,并维持细胞种的延续。培养的细胞形成单层汇合以后,由于密度过大生存空间不足而引起营养枯竭,将培养的细胞分散,从容器中取出,以1:2或1:3以上的比率转移到另外的容器中进行培养,即为传代培养;细胞“一代”指从细胞接种到分离再培养的一段期间,与细胞世代或倍增不同。在一代中,细胞培增3~6次。细胞传代后,一般经过三个阶段:游离期、指数增生期和停止期。常用细胞分裂指数表示细胞增殖的旺盛程度,即细胞群的分裂相数/100个细胞。一般细胞分裂指数介于0.2%~0.5%,肿瘤细胞可达3~5%;细胞接种2~3天分裂增殖旺盛,是活力ZuiHAO时期,称指数增生期(对数生长期),适宜进行各种试验。实验步骤:1.将长成的培养细胞从二氧化碳培养箱中取出,在超净工作台中倒掉瓶内的培养,加入少许消化。(以面盖住细胞为宜),静置5~10分钟。2.在倒置镜下观察被消化的细胞,如果细胞变圆,相互之间不再连接成片,这时应立即在超净台中将消化倒掉,加入3~5ml新鲜培养,吹打,制成细胞悬。3.将细胞悬吸出2ml左右,加到另一个培养瓶中并向每个瓶中分别加3ml左右培养,盖HAO瓶塞,送回二氧化碳培养箱中,继续进行培养。一般情况,传代后的细胞在2小时左右就能附着在培养瓶壁上,2~4天就可在瓶内形成单层,需要再次进行传代。
COLO 684 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:悬浮生长;形态特性:详见产品说明书;相关产品有:NCI-SNU-620细胞、AtT-20细胞、SKMEL2细胞
SK-N-BE(1) Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:AZ-521细胞、NOR-10细胞、RAMOS2G64C10细胞
Neukoplast Cells;背景说明:NK细胞;淋巴瘤;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:HKF细胞、Laboratory of Allergic Diseases 2细胞、TOV21G细胞
P3X63Ag8653 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:A704细胞、CHO cell clone K1细胞、HEC-1B细胞
SK-ES-1 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:5传代;每周换液2-3次;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:上皮样;相关产品有:CD18/HPAF细胞、GCT细胞、N87细胞
Me-Wo Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:5传代,2-3天换液1次。;生长特性:混合生长;形态特性:成纤维细胞;相关产品有:CEM细胞、HO1-N-1细胞、SW837细胞
B16BL6 Cells;背景说明:黑色素瘤;雄性;C57BL/6;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:HEI193细胞、Rat Basophilic Leukemia-1细胞、MSF细胞
Mc Coy Cells;背景说明:成纤维 Cells;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:FHs74 Int细胞、526细胞、KASUMI1细胞
KU-812 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代。3天内可长满;生长特性:悬浮生长;形态特性:骨髓母细胞;相关产品有:PC14细胞、2008细胞、NKT细胞
2780 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:aTC1 Clone 6细胞、T-47D细胞、HCC-1428细胞
WM115F Cells;背景说明:黑色素瘤;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:WIL2S细胞、KPL-1细胞、LLC PK1细胞
NR8383.1 Cells;背景说明:NR8383(正常大鼠,1983年8月3日)来源于肺灌洗时的正常大鼠肺泡巨噬细胞。细胞在gerbil肺细胞连续培养液存在下培养了大约8-9个月。随后,不再需要外源生长因子。通过有限稀释法从单个细胞克隆并亚克隆NR8383细胞,并三次用软琼脂亚克隆。细胞表现出巨噬细胞的特性,吞噬酵母多糖和铜绿,非特异性脂酶活性,Fc受体,氧化降解;分泌IL-1,TNFbeta和IL-6,可重复地响应外源生长因子。NR8383细胞响应博莱霉素,分泌TGFbeta前体。在博莱霉素刺激下,TGFbe;传代方法:1:2传代;生长特性:半贴壁生长;形态特性:巨噬细胞;相关产品有:NCIH520细胞、COC1/CDDP细胞、A2058细胞
TOV112D Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代,3-4天换液1次。;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:MKN7细胞、EB1细胞、HIC细胞
RWPE-1 Cells;背景说明:肿瘤抑制基因: p53 + [PubMed: 9214605] pRB + [PubMed: 9214605] 一位正常男性前列腺组织切片的周围区域的上皮细胞用单拷贝的人乳头瘤病毒的18(HPV-18)进行转化,建立了RWPE-1 (ATCC CRL-11609) 细胞株 [PubMed: 9214605]. 在三维Matrigel培养时,在雄激素作用下,RWPE-1细胞形成腺胞并向培养基中分泌PSA。[PubMed: 11170142]. 当与Matrigel或基质细胞混合注射雄性裸鼠时,RWPE-1细胞也能形成腺胞[PubMed: 11304724] 并产生PSA。 来源于RWPE-1的细胞再用Kirstin鼠类肿瘤病毒转染Ki-ras基因,建立了能成瘤的RWPE-2细胞株(ATCC CRL-11610) [PubMed: 9214605] 和 RWPE2-W99 (ATCC CRL-2853) 细胞株。 另外,用N--N-(MNU)处理RWPE-1,建立了一系列模拟前列腺癌进程中不同时期的成瘤细胞株。 它们是 WPE1-NA22 (ATCC CRL-2849), WPE1-NB14 (ATCC CRL-2850, WPE1-NB11 (ATCC CRL-2851) 和 WPE1-NB26 (ATCC CRL-2852) 细胞株。 据提供者报道,RWPE-1 细胞株(ATCC CRL-11609)经过检测,乙肝、丙肝、人免疫缺陷病毒都呈阴性。;传代方法:1:3传代,2-3天传一代。;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:NCI-H2073细胞、EFM192A细胞、TE3细胞
X63 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:MPASMC细胞、C8-D1A细胞、SU4细胞
VMCUB-1 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Huh 7.5.1细胞、NB1RGB细胞、NOR10细胞
NCIH1623 Cells;背景说明:详见相关文献介绍;传代方法:每周换液2-3次。;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:MRC5细胞、PLA801-95D细胞、HRA19细胞
Abcam HEK293T EEF1B2 KO Cells(提供STR鉴定图谱)
AG09971 Cells(提供STR鉴定图谱)
BayGenomics ES cell line NPX366 Cells(提供STR鉴定图谱)
BayGenomics ES cell line XC588 Cells(提供STR鉴定图谱)
BT-IMF Cells(提供STR鉴定图谱)
COSB Cells(提供STR鉴定图谱)
DA04058 Cells(提供STR鉴定图谱)
DA05762 Cells(提供STR鉴定图谱)
GM00008 Cells(提供STR鉴定图谱)
NCI-H125 Cells;背景说明:腺鳞状肺癌;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:J82COT细胞、SKMEL28细胞、HT144-mel细胞
T98-G Cells;背景说明:详见相关文献介绍;传代方法:按1:3传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:H-28细胞、D341 Med细胞、OV1/P细胞
HCC0366 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:MCF/Adr细胞、RN-c细胞、NTC200细胞
NCIH524 Cells;背景说明:该细胞1982年建系,源自非小细胞肺癌男性患者的转移淋巴结。;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮生长;形态特性:圆形细胞;相关产品有:SDBMSC细胞、MDA-415细胞、Kit225细胞
PVEC Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:L-5178-Y细胞、MDAMB453细胞、HeLaS3细胞
Colo741 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:KYSE-410细胞、EnCa1细胞、TE671细胞
HBVP Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:NCIH1793细胞、Panc-8_13细胞、Reuber H35细胞
NIH:OVCAR-3 Cells;背景说明:该细胞1982年由T.C. Hamilton等建系,源自一位60卵巢腺癌的腹水,是卵巢癌抗药性研究的模型。;传代方法:1:2—1:4传代,每周换液2—3次;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:Super Tube细胞、HDLM-2细胞、IMR32细胞
SK-OV-3人卵巢癌细胞全年复苏|已有STR图谱
Panc_02_03 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:OCI-LY-19细胞、HCC1954-BL细胞、KU1919细胞
C-6 Cells;背景说明:胶质细胞株C6是由Benda等用N-nitrosomethylurea诱导的大鼠胶质瘤克隆,并经过一系列的体外培养和动物传代交替后建成的。 当细胞从低密度生长到满瓶时,S-100产量增加10倍。;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:MRC9细胞、JIMT细胞、GM-637细胞
WM2664 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:A-172 MG细胞、P3J HR1-K细胞、KYSE 50细胞
H-774 Cells;背景说明:详见相关文献介绍;传代方法:每周换液2次。;生长特性:悬浮生长;形态特性:详见产品说明书;相关产品有:GM00637细胞、SKNBE细胞、GB-1细胞
NKM1 Cells;背景说明:急性髓系白血病;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:ECC 10细胞、RD-2细胞、766T细胞
EFO 27 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:TU 686细胞、Kit-225细胞、JB6 [Mouse]细胞
NCI-H2342 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:6传代 ;生长特性:贴壁生长;形态特性:上皮样;相关产品有:174xCEM.T2细胞、NCIH889细胞、hEEC细胞
H3730 Cells(提供STR鉴定图谱)
HAP1 RAB39B (-) 1 Cells(提供STR鉴定图谱)
BEL/FU Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:KOSC-2 cl3-43细胞、HBL1细胞、RT-BM细胞
NCI H929 Cells;背景说明:详见相关文献介绍;传代方法:保持细胞密度在5×105—1×106 cells/ml之间,每周换液2—3次;生长特性:悬浮生长 ;形态特性:淋巴母细胞样;相关产品有:SUM 159细胞、J82COT细胞、CCRF-CEM细胞
B 95.8 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:NS20Y细胞、KBM5细胞、HFE-145细胞
BEAS-2B Cells;背景说明:从一位非癌个体的正常人支气管上皮病理切片分离出上皮细胞。这些细胞用腺病毒12-SV40病毒杂交病毒感染并克隆。DEAS-2B细胞保留了对血清反应进行鳞关分化的能力,并有用于筛选诱导或影响分化及致癌的化学或生物制剂。细胞角蛋白及SV40抗原染色阳性。;传代方法:消化3-5分钟。1:2。3天内可长满;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:PA-1细胞、143TK-细胞、HCC0078细胞
NPA-87 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:RC92A细胞、H-4细胞、SUPT-1细胞
Astrocyte type I clone Cells;背景说明:该永生化细胞系源自出生8天小鼠小脑组织,由B Pessac, D Trisler建立。该细胞具有小神经胶质细胞特征。该细胞为GFAP阳性细胞,除此之外,没有检测到其它神经胶质神经元或小神经胶质细胞的分子标记。;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:NCI-SNU-1细胞、H2.35细胞、SHI1细胞
MRC-V Cells;背景说明:MRC-5细胞系来自14周龄男性胎儿的正常肺组织,该细胞老化前能传代42~46个倍增时间。;传代方法:1:2-1:5传代;每周1-2次。;生长特性:贴壁生长;形态特性:成纤维细胞样;相关产品有:Human Pancreatic Duct Epithelial细胞、HSC 3细胞、HO8910细胞
SNUC2A Cells;背景说明:详见相关文献介绍;传代方法:每周两次换液;生长特性:松散附着、多单元的聚合;形态特性:上皮细胞样;相关产品有:A 549细胞、Panc_02_03细胞、Michigan Cancer Foundation-12A细胞
HuH7-SLC51A-KO-c1 Cells(提供STR鉴定图谱)
L5178Y BLM-A2-R Cells(提供STR鉴定图谱)
MT58 Cells(提供STR鉴定图谱)
OMC-6 Cells(提供STR鉴定图谱)
Royan D8 Cells(提供STR鉴定图谱)
UCSD083i-40-2 Cells(提供STR鉴定图谱)
ZR-75-1 XI-36 Cells(提供STR鉴定图谱)
HEV0146 Cells(提供STR鉴定图谱)
CC-LP-I Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:MES-SA Dx5细胞、NCIH2052细胞、OVHM细胞
FC33 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:MonoMac1细胞、HEK-A细胞、AM38细胞
MLE 12 Cells;背景说明:肺;上皮细胞;SV40转化;FVB/N;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:MLA144细胞、MDA MB 453细胞、GM03569细胞
PIEC Cells;背景说明:髋动脉;内皮 Cells;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:SK-SH-SY5Y细胞、GC-1 spg细胞、WiDr-TC细胞
Panc02 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:P815细胞、KTA-7细胞、CHOS细胞
Panc02 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:P815细胞、KTA-7细胞、CHOS细胞
WM 239A Cells;背景说明:黑色素瘤;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:WEHI 164 TC细胞、NCI-H1672细胞、JHH-2细胞
H-441 Cells;背景说明:NCI-H441建系于1982年(A.F.Gazdar,etal.)。该细胞分离自一名肺腺癌患者的心包液。该细胞能在半固体琼脂糖中形成克隆,并能表达肺泡表面活性蛋白A。该细胞在有血清培养基中倍增时间为58小时,在无血清培养基中倍增时间为99-138小时。;传代方法:1:3传代,2-3天传一代;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:769-P细胞、OVCAR 5细胞、SUM-190PT细胞
NCI-H226 Cells;背景说明:1980年分离建立。;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:Doubling time: ~50 hours (ATCC).细胞、H820细胞、RA细胞
NCIH1048 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:8传代;;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:SW-1573细胞、OCIAML5细胞、MDCC-MSB1细胞
PANC-02-03 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:HEK-293-EBNA细胞、MSTO211H细胞、MeWo细胞
Panc02-H0 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:PANC-08-13细胞、Y-1细胞、VA-ES-BJ细胞
SK-GT-2 Cells;背景说明:胃底癌;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:D-407细胞、KP-N-RT-BM-1细胞、H-774细胞
SKCO 1 Cells;背景说明:该细胞来源于结直肠病人的转移性腹水。;传代方法:1:2-1:3传代,每周2-3次。;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:TE32细胞、JJN3细胞、MDA-MB 231细胞
NCIH2227 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代;每周换液2次。;生长特性:该细胞既有悬浮生长,又有贴壁生长;形态特性:上皮细胞;相关产品有:OK细胞、RCC-JF细胞、HMrSV5细胞
T24/DDP10 Cells(提供STR鉴定图谱)
GM637A Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:HO-8910 PM细胞、686LN-M4e细胞、Hs 746.T细胞
NCI-H2085 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代 ;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:L-6细胞、H-7721细胞、HT55细胞
NCI-H1648 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:CT26-clone 25细胞、UPCI:SCC154细胞、MSTO-211H细胞
CMT.64 Cells;背景说明:肺腺癌;雌性;C57;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:Mv.1.Lu细胞、NCI-H2135细胞、Mel-RM细胞
NCI-HUT-520 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代;2-3天换液1次;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:P3J HR-1细胞、RS(4;11)细胞、MF2059细胞
Lewis lung carcinoma line 1 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:MDA157细胞、IPEC-1细胞、SNU119细胞
Pt K1 (NBL-3) Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Jurkat 77细胞、Hs-281-T细胞、Huh-7.5.1细胞
149 PT Cells;背景说明:乳腺癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:Hs 578T细胞、H-9细胞、SNU878细胞
SK-OV-3人卵巢癌细胞全年复苏|已有STR图谱
G-292, clone A141B1 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Hs 766.T细胞、SCC-1395细胞、MT4细胞
GM03569B Cells;背景说明:该细胞是由绵羊红细胞免疫的BALB/c小鼠脾细胞和P3X63Ag8骨髓瘤细胞融合得到的。该细胞不分泌免疫球蛋白,对20μg/ml的8-氮鸟嘌呤有抗性,对HAT比较敏感;该细胞可以作为细胞融合时的B细胞组分用于制备杂交瘤;鼠痘病毒阴性。;传代方法:1:2传代;生长特性:悬浮生长;形态特性:淋巴母细胞样;圆形;相关产品有:NCI-H1819细胞、NUGC-2细胞、EBC-1/original细胞
NCIH1436 Cells;背景说明:详见相关文献介绍;传代方法:随细胞的密度而增加;生长特性:悬浮生长;形态特性:详见产品说明书;相关产品有:RD-ES细胞、CAMA1细胞、Renal Carcinoma细胞
MC/CAR Cells;背景说明:B淋巴;EBV转化;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:A-2058细胞、ECGI10细胞、16-HBE14o细胞
TI-73 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代;每周换液2-3次。;生长特性:贴壁生长;形态特性:成纤维细胞;相关产品有:MM1细胞、SNU449细胞、NK-92 transfected with MFG-hIL2细胞
NCI-SNU-886 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Murine Long bone Osteocyte-Y4细胞、KYSE 140细胞、BNL CL2细胞
BayGenomics ES cell line RRM081 Cells(提供STR鉴定图谱)
BayGenomics ES cell line XN619 Cells(提供STR鉴定图谱)
GSI-1 Cells(提供STR鉴定图谱)
OX-36 clone 3 Cells(提供STR鉴定图谱)
3Y1tsE204 Cells(提供STR鉴定图谱)
MDA-MB-468/Cas9-hyg Cells(提供STR鉴定图谱)
" "PubMed=6582512; DOI=10.1073/pnas.81.2.568; PMCID=PMC344720
Mattes M.J., Cordon-Cardo C., Lewis J.L. Jr., Old L.J., Lloyd K.O.
Cell surface antigens of human ovarian and endometrial carcinoma defined by mouse monoclonal antibodies.
Proc. Natl. Acad. Sci. U.S.A. 81:568-572(1984)
PubMed=4016745
Buick R.N., Pullano R., Trent J.M.
Comparative properties of five human ovarian adenocarcinoma cell lines.
Cancer Res. 45:3668-3676(1985)
PubMed=3518877; DOI=10.3109/07357908609038260
Fogh J.
Human tumor lines for cancer research.
Cancer Invest. 4:157-184(1986)
PubMed=3804493; DOI=10.1002/ijc.2910390216
Hill B.T., Whelan R.D.H., Gibby E.M., Sheer D., Hosking L.K., Shellard S.A., Rupniak H.T.
Establishment and characterisation of three new human ovarian carcinoma cell lines and initial evaluation of their potential in experimental chemotherapy studies.
Int. J. Cancer 39:219-225(1987)
PubMed=3335022
Alley M.C., Scudiero D.A., Monks A., Hursey M.L., Czerwinski M.J., Fine D.L., Abbott B.J., Mayo J.G., Shoemaker R.H., Boyd M.R.
Feasibility of drug screening with panels of human tumor cell lines using a microculture tetrazolium assay.
Cancer Res. 48:589-601(1988)
PubMed=2653399; DOI=10.1038/bjc.1989.108; PMCID=PMC2247140
Hills C.A., Kelland L.R., Abel G., Siracky J., Wilson A.P., Harrap K.R.
Biological properties of ten human ovarian carcinoma cell lines: calibration in vitro against four platinum complexes.
Br. J. Cancer 59:527-534(1989)
PubMed=1892748; DOI=10.1038/bjc.1991.279; PMCID=PMC1977519
Mistry P., Kelland L.R., Abel G., Sidhar S., Harrap K.R.
The relationships between glutathione, glutathione-S-transferase and cytotoxicity of platinum drugs and melphalan in eight human ovarian carcinoma cell lines.
Br. J. Cancer 64:215-220(1991)
PubMed=2041050; DOI=10.1093/jnci/83.11.757
Monks A., Scudiero D.A., Skehan P., Shoemaker R.H., Paull K.D., Vistica D.T., Hose C.D., Langley J., Cronise P., Vaigro-Wolff A., Gray-Goodrich M., Campbell H., Mayo J.G., Boyd M.R.
Feasibility of a high-flux anticancer drug screen using a diverse panel of cultured human tumor cell lines.
J. Natl. Cancer Inst. 83:757-766(1991)
PubMed=1423261
Mistry P., Kelland L.R., Loh S.Y., Abel G., Murrer B.A., Harrap K.R.
Comparison of cellular accumulation and cytotoxicity of cisplatin with that of tetraplatin and amminedibutyratodichloro(cyclohexylamine)platinum(IV) (JM221) in human ovarian carcinoma cell lines.
Cancer Res. 52:6188-6193(1992)
PubMed=7718330; DOI=10.1016/0959-8049(94)00472-H
Baguley B.C., Marshall E.S., Whittaker J.R., Dotchin M.C., Nixon J., McCrystal M.R., Finlay G.J., Matthews J.H.L., Holdaway K.M., van Zijl P.L.
Resistance mechanisms determining the in vitro sensitivity to paclitaxel of tumour cells cultured from patients with ovarian cancer.
Eur. J. Cancer 31A:230-237(1995)
PubMed=8557231; DOI=10.1006/gyno.1996.0014
Skilling J.S., Squatrito R.C., Connor J.P., Niemann T., Buller R.E.
p53 gene mutation analysis and antisense-mediated growth inhibition of human ovarian carcinoma cell lines.
Gynecol. Oncol. 60:72-80(1996)
PubMed=9041185
Johnson S.W., Laub P.B., Beesley J.S., Ozols R.F., Hamilton T.C.
Increased platinum-DNA damage tolerance is associated with cisplatin resistance and cross-resistance to various chemotherapeutic agents in unrelated human ovarian cancer cell lines.
Cancer Res. 57:850-856(1997)
PubMed=9554442; DOI=10.1093/jnci/90.8.597
Sabichi A.L., Hendricks D.T., Bober M.A., Birrer M.J.
Retinoic acid receptor beta expression and growth inhibition of gynecologic cancer cells by the synthetic retinoid N-(4-hydroxyphenyl) retinamide.
J. Natl. Cancer Inst. 90:597-605(1998)
PubMed=9698466; DOI=10.1006/gyno.1998.5039
Maxwell G.L., Risinger J.I., Tong B., Shaw H., Barrett J.C., Berchuck A., Futreal P.A.
Mutation of the PTEN tumor suppressor gene is not a feature of ovarian cancers.
Gynecol. Oncol. 70:13-16(1998)
PubMed=10318951; DOI=10.1073/pnas.96.10.5722; PMCID=PMC21927
Lau K.-M., Mok S.C., Ho S.-M.
Expression of human estrogen receptor-alpha and -beta, progesterone receptor, and androgen receptor mRNA in normal and malignant ovarian epithelial cells.
Proc. Natl. Acad. Sci. U.S.A. 96:5722-5727(1999)
PubMed=10700174; DOI=10.1038/73432
Ross D.T., Scherf U., Eisen M.B., Perou C.M., Rees C., Spellman P.T., Iyer V.R., Jeffrey S.S., van de Rijn M., Waltham M.C., Pergamenschikov A., Lee J.C.F., Lashkari D., Shalon D., Myers T.G., Weinstein J.N., Botstein D., Brown P.O.
Systematic variation in gene expression patterns in human cancer cell lines.
Nat. Genet. 24:227-235(2000)
PubMed=10972993; DOI=10.1002/1098-2744(200008)28:4<236::aid-mc6>3.0.CO;2-H
Rauh-Adelmann C., Lau K.-M., Sabeti N., Long J.P., Mok S.C., Ho S.-M.
Altered expression of BRCA1, BRCA2, and a newly identified BRCA2 exon 12 deletion variant in malignant human ovarian, prostate, and breast cancer cell lines.
Mol. Carcinog. 28:236-246(2000)
PubMed=11416159; DOI=10.1073/pnas.121616198; PMCID=PMC35459
Masters J.R.W., Thomson J.A., Daly-Burns B., Reid Y.A., Dirks W.G., Packer P., Toji L.H., Ohno T., Tanabe H., Arlett C.F., Kelland L.R., Harrison M., Virmani A.K., Ward T.H., Ayres K.L., Debenham P.G.
Short tandem repeat profiling provides an international reference standard for human cell lines.
Proc. Natl. Acad. Sci. U.S.A. 98:8012-8017(2001)
PubMed=11565033; DOI=10.1038/35095068
Moberg K.H., Bell D.W., Pronk-Wahrer D.C.R., Haber D.A., Hariharan I.K.
Archipelago regulates cyclin E levels in Drosophila and is mutated in human cancer cell lines.
Nature 413:311-316(2001)
PubMed=11789735; DOI=10.1309/4NCM-QJ9W-QM0J-6QJE
Rhodes A., Jasani B., Couturier J., McKinley M.J., Morgan J.M., Dodson A.R., Navabi H., Miller K.D., Balaton A.J.
A formalin-fixed, paraffin-processed cell line standard for quality control of immunohistochemical assay of HER-2/neu expression in breast cancer.
Am. J. Clin. Pathol. 117:81-89(2002)
PubMed=11793438; DOI=10.1002/gcc.1221
Rao P.H., Harris C.P., Lu X.-Y., Li X.-N., Mok S.C., Lau C.C.
Multicolor spectral karyotyping of serous ovarian adenocarcinoma.
Genes Chromosomes Cancer 33:123-132(2002)
PubMed=12080474; DOI=10.1038/sj.onc.1205542
Manzano R.G., Montuenga L.M., Dayton M.A., Dent P., Kinoshita I., Vicent S., Gardner G.J., Nguyen P., Choi Y.-H., Trepel J.B., Auersperg N., Birrer M.J.
CL100 expression is down-regulated in advanced epithelial ovarian cancer and its re-expression decreases its malignant potential.
Oncogene 21:4435-4447(2002)
PubMed=12417041; DOI=10.1111/j.1349-7006.2002.tb01213.x; PMCID=PMC5926887
Watanabe T., Imoto I., Katahira T., Hirasawa A., Ishiwata I., Emi M., Takayama M., Sato A., Inazawa J.
Differentially regulated genes as putative targets of amplifications at 20q in ovarian cancers.
Jpn. J. Cancer Res. 93:1114-1122(2002)
PubMed=12960427; DOI=10.1091/mbc.E03-05-0279; PMCID=PMC266758
Schaner M.E., Ross D.T., Ciaravino G., Sorlie T., Troyanskaya O.G., Diehn M., Wang Y.C., Duran G.E., Sikic T.L., Caldeira S., Skomedal H., Tu I.-P., Hernandez-Boussard T., Johnson S.W., O'Dwyer P.J., Fero M.J., Kristensen G.B., Borresen-Dale A.-L., Hastie T.J., Tibshirani R., van de Rijn M., Teng N.N.H., Longacre T.A., Botstein D., Brown P.O., Sikic B.I.
Gene expression patterns in ovarian carcinomas.
Mol. Biol. Cell 14:4376-4386(2003)
PubMed=15748285; DOI=10.1186/1479-5876-3-11; PMCID=PMC555742
Adams S., Robbins F.-M., Chen D., Wagage D., Holbeck S.L., Morse H.C. 3rd, Stroncek D., Marincola F.M.
HLA class I and II genotype of the NCI-60 cell lines.
J. Transl. Med. 3:11.1-11.8(2005)
PubMed=16380993; DOI=10.1002/ijc.21671
Huang K.-C., Park D.C., Ng S.-K., Lee J.Y., Ni X.-Y., Ng W.-C., Bandera C.A., Welch W.R., Berkowitz R.S., Mok S.C., Ng S.-W.
Selenium binding protein 1 in ovarian cancer.
Int. J. Cancer 118:2433-2440(2006)
PubMed=17088437; DOI=10.1158/1535-7163.MCT-06-0433; PMCID=PMC2705832
Ikediobi O.N., Davies H.R., Bignell G.R., Edkins S., Stevens C., O'Meara S., Santarius T., Avis T., Barthorpe S., Brackenbury L., Buck G., Butler A.P., Clements J., Cole J., Dicks E., Forbes S., Gray K., Halliday K., Harrison R., Hills K., Hinton J., Hunter C., Jenkinson A., Jones D., Kosmidou V., Lugg R., Menzies A., Miroo T., Parker A., Perry J., Raine K.M., Richardson D., Shepherd R., Small A., Smith R., Solomon H., Stephens P.J., Teague J.W., Tofts C., Varian J., Webb T., West S., Widaa S., Yates A., Reinhold W.C., Weinstein J.N., Stratton M.R., Futreal P.A., Wooster R.
Mutation analysis of 24 known cancer genes in the NCI-60 cell line set.
Mol. Cancer Ther. 5:2606-2612(2006)
PubMed=17726699; DOI=10.1002/gcc.20492
Buys T.P.H., Chari R., Lee E.H.-L., Zhang M., MacAulay C., Lam S., Lam W.L., Ling V.
Genetic changes in the evolution of multidrug resistance for cultured human ovarian cancer cells.
Genes Chromosomes Cancer 46:1069-1079(2007)
PubMed=19372543; DOI=10.1158/1535-7163.MCT-08-0921; PMCID=PMC4020356
Lorenzi P.L., Reinhold W.C., Varma S., Hutchinson A.A., Pommier Y., Chanock S.J., Weinstein J.N.
DNA fingerprinting of the NCI-60 cell line panel.
Mol. Cancer Ther. 8:713-724(2009)
PubMed=20164919; DOI=10.1038/nature08768; PMCID=PMC3145113
Bignell G.R., Greenman C.D., Davies H.R., Butler A.P., Edkins S., Andrews J.M., Buck G., Chen L., Beare D., Latimer C., Widaa S., Hinton J., Fahey C., Fu B.-Y., Swamy S., Dalgliesh G.L., Teh B.T., Deloukas P., Yang F.-T., Campbell P.J., Futreal P.A., Stratton M.R.
Signatures of mutation and selection in the cancer genome.
Nature 463:893-898(2010)
PubMed=20204287; DOI=10.3892/or_00000728; PMCID=PMC2909445
Langland G.T., Yannone S.M., Langland R.A., Nakao A., Guan Y.-H., Long S.B.T., Vonguyen L., Chen D.J., Gray J.W., Chen F.-Q.
Radiosensitivity profiles from a panel of ovarian cancer cell lines exhibiting genetic alterations in p53 and disparate DNA-dependent protein kinase activities.
Oncol. Rep. 23:1021-1026(2010)
PubMed=20215515; DOI=10.1158/0008-5472.CAN-09-3458; PMCID=PMC2881662
Rothenberg S.M., Mohapatra G., Rivera M.N., Winokur D., Greninger P., Nitta M., Sadow P.M., Sooriyakumar G., Brannigan B.W., Ulman M.J., Perera R.M., Wang R., Tam A., Ma X.-J., Erlander M., Sgroi D.C., Rocco J.W., Lingen M.W., Cohen E.E.W., Louis D.N., Settleman J., Haber D.A.
A genome-wide screen for microdeletions reveals disruption of polarity complex genes in diverse human cancers.
Cancer Res. 70:2158-2164(2010)
PubMed=22068913; DOI=10.1073/pnas.1111840108; PMCID=PMC3219108
Gillet J.-P., Calcagno A.M., Varma S., Marino M., Green L.J., Vora M.I., Patel C., Orina J.N., Eliseeva T.A., Singal V., Padmanabhan R., Davidson B., Ganapathi R., Sood A.K., Rueda B.R., Ambudkar S.V., Gottesman M.M.
Redefining the relevance of established cancer cell lines to the study of mechanisms of clinical anti-cancer drug resistance.
Proc. Natl. Acad. Sci. U.S.A. 108:18708-18713(2011)
PubMed=21912889; DOI=10.1007/s10637-011-9744-z
Sutherland H.S., Hwang I.Y., Marshall E.S., Lindsay B.S., Denny W.A., Gilchrist C., Joseph W.R., Greenhalgh D., Richardson E., Kestell P., Ding A., Baguley B.C.
Therapeutic reactivation of mutant p53 protein by quinazoline derivatives.
Invest. New Drugs 30:2035-2045(2012)
PubMed=22328975; DOI=10.1158/2159-8290.CD-11-0170; PMCID=PMC3274821
Hanrahan A.J., Schultz N., Westfal M.L., Sakr R.A., Giri D.D., Scarperi S., Janakiraman M., Olvera N., Stevens E.V., She Q.-B., Aghajanian C., King T.A., de Stanchina E., Spriggs D.R., Heguy A., Taylor B.S., Sander C., Rosen N., Levine D.A., Solit D.B.
Genomic complexity and AKT dependence in serous ovarian cancer.
Cancer Discov. 2:56-67(2012)
PubMed=22336246; DOI=10.1016/j.bmc.2012.01.017
Kong D.-X., Yamori T.
JFCR39, a panel of 39 human cancer cell lines, and its application in the discovery and development of anticancer drugs.
Bioorg. Med. Chem. 20:1947-1951(2012)
PubMed=22347499; DOI=10.1371/journal.pone.0031628; PMCID=PMC3276511
Ruan X.-Y., Kocher J.-P.A., Pommier Y., Liu H.-F., Reinhold W.C.
Mass homozygotes accumulation in the NCI-60 cancer cell lines as compared to HapMap trios, and relation to fragile site location.
PLoS ONE 7:E31628-E31628(2012)
PubMed=22384151; DOI=10.1371/journal.pone.0032096; PMCID=PMC3285665
Lee J.-S., Kim Y.K., Kim H.J., Hajar S., Tan Y.L., Kang N.-Y., Ng S.H., Yoon C.N., Chang Y.-T.
Identification of cancer cell-line origins using fluorescence image-based phenomic screening.
PLoS ONE 7:E32096-E32096(2012)
PubMed=22460905; DOI=10.1038/nature11003; PMCID=PMC3320027
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=22585861; DOI=10.1158/2159-8290.CD-11-0224; PMCID=PMC5057396
Marcotte R., Brown K.R., Suarez Saiz F.J., Sayad A., Karamboulas K., Krzyzanowski P.M., Sircoulomb F., Medrano M., Fedyshyn Y., Koh J.L.-Y., van Dyk D., Fedyshyn B., Luhova M., Brito G.C., Vizeacoumar F.J., Vizeacoumar F.S., Datti A., Kasimer D., Buzina A., Mero P., Misquitta C., Normand J., Haider M., Ketela T., Wrana J.L., Rottapel R., Neel B.G., Moffat J.
Essential gene profiles in breast, pancreatic, and ovarian cancer cells.
Cancer Discov. 2:172-189(2012)
PubMed=22628656; DOI=10.1126/science.1218595; PMCID=PMC3526189
Jain M., Nilsson R., Sharma S., Madhusudhan N., Kitami T., Souza A.L., Kafri R., Kirschner M.W., Clish C.B., Mootha V.K.
Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation.
Science 336:1040-1044(2012)
PubMed=22705003; DOI=10.1016/j.humpath.2012.03.011
Rahman M., Nakayama K., Rahman M.T., Nakayama N., Ishikawa M., Katagiri A., Iida K., Nakayama S., Otsuki Y., Shih I.-M., Miyazaki K.
Clinicopathologic and biological analysis of PIK3CA mutation in ovarian clear cell carcinoma.
Hum. Pathol. 43:2197-2206(2012)
PubMed=22710073; DOI=10.1016/j.ygyno.2012.06.017; PMCID=PMC3432677
Korch C.T., Spillman M.A., Jackson T.A., Jacobsen B.M., Murphy S.K., Lessey B.A., Jordan V.C., Bradford A.P.
DNA profiling analysis of endometrial and ovarian cell lines reveals misidentification, redundancy and contamination.
Gynecol. Oncol. 127:241-248(2012)
PubMed=23415752; DOI=10.1016/j.molonc.2012.12.007; PMCID=PMC4106023
Stordal B., Timms K., Farrelly A., Gallagher D., Busschots S., Renaud M., Thery J., Williams D., Potter J., Tran T., Korpanty G., Cremona M., Carey M.S., Li J., Li Y., Aslan O., O'Leary J.J., Mills G.B., Hennessy B.T.
BRCA1/2 mutation analysis in 41 ovarian cell lines reveals only one functionally deleterious BRCA1 mutation.
Mol. Oncol. 7:567-579(2013)
PubMed=23839242; DOI=10.1038/ncomms3126; PMCID=PMC3715866
Domcke S., Sinha R., Levine D.A., Sander C., Schultz N.
Evaluating cell lines as tumour models by comparison of genomic profiles.
Nat. Commun. 4:2126.1-2126.10(2013)
PubMed=23856246; DOI=10.1158/0008-5472.CAN-12-3342; PMCID=PMC4893961
Abaan O.D., Polley E.C., Davis S.R., Zhu Y.-L.J., Bilke S., Walker R.L., Pineda M.A., Gindin Y., Jiang Y., Reinhold W.C., Holbeck S.L., Simon R.M., Doroshow J.H., Pommier Y., Meltzer P.S.
The exomes of the NCI-60 panel: a genomic resource for cancer biology and systems pharmacology.
Cancer Res. 73:4372-4382(2013)
PubMed=23933261; DOI=10.1016/j.celrep.2013.07.018
Moghaddas Gholami A., Hahne H., Wu Z.-X., Auer F.J., Meng C., Wilhelm M., Kuster B.
Global proteome analysis of the NCI-60 cell line panel.
Cell Rep. 4:609-620(2013)
PubMed=24023729; DOI=10.1371/journal.pone.0072162; PMCID=PMC3762837
Anglesio M.S., Wiegand K.C., Melnyk N., Chow C., Salamanca C.M., Prentice L.M., Senz J., Yang W., Spillman M.A., Cochrane D.R., Shumansky K., Shah S.P., Kalloger S.E., Huntsman D.G.
Type-specific cell line models for type-specific ovarian cancer research.
PLoS ONE 8:E72162-E72162(2013)
PubMed=24279929; DOI=10.1186/2049-3002-1-20; PMCID=PMC4178206
Dolfi S.C., Chan L.L.-Y., Qiu J., Tedeschi P.M., Bertino J.R., Hirshfield K.M., Oltvai Z.N., Vazquez A.
The metabolic demands of cancer cells are coupled to their size and protein synthesis rates.
Cancer Metab. 1:20.1-20.13(2013)
PubMed=24434149; DOI=10.1016/j.bbrc.2013.12.070
Sinha A., Ignatchenko V., Ignatchenko A., Mejia-Guerrero S., Kislinger T.
In-depth proteomic analyses of ovarian cancer cell line exosomes reveals differential enrichment of functional categories compared to the NCI 60 proteome.
Biochem. Biophys. Res. Commun. 445:694-701(2014)
PubMed=24670534; DOI=10.1371/journal.pone.0092047; PMCID=PMC3966786
Varma S., Pommier Y., Sunshine M., Weinstein J.N., Reinhold W.C.
High resolution copy number variation data in the NCI-60 cancer cell lines from whole genome microarrays accessible through CellMiner.
PLoS ONE 9:E92047-E92047(2014)
PubMed=25230021; DOI=10.1371/journal.pone.0103988; PMCID=PMC4167545
Beaufort C.M., Helmijr J.C.A., Piskorz A.M., Hoogstraat M., Ruigrok-Ritstier K., Besselink N., Murtaza M., van IJcken W.F.J., Heine A.A.J., Smid M., Koudijs M.J., Brenton J.D., Berns E.M.J.J., Helleman J.
Ovarian cancer cell line panel (OCCP): clinical importance of in vitro morphological subtypes.
PLoS ONE 9:E103988-E103988(2014)
PubMed=25984343; DOI=10.1038/sdata.2014.35; PMCID=PMC4432652
Cowley G.S., Weir B.A., Vazquez F., Tamayo P., Scott J.A., Rusin S., East-Seletsky A., Ali L.D., Gerath W.F.J., Pantel S.E., Lizotte P.H., Jiang G.-Z., Hsiao J., Tsherniak A., Dwinell E., Aoyama S., Okamoto M., Harrington W., Gelfand E.T., Green T.M., Tomko M.J., Gopal S., Wong T.C., Li H.-B., Howell S., Stransky N., Liefeld T., Jang D., Bistline J., Meyers B.H., Armstrong S.A., Anderson K.C., Stegmaier K., Reich M., Pellman D., Boehm J.S., Mesirov J.P., Golub T.R., Root D.E., Hahn W.C.
Parallel genome-scale loss of function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies.
Sci. Data 1:140035-140035(2014)
PubMed=25485619; DOI=10.1038/nbt.3080
Klijn C., Durinck S., Stawiski E.W., Haverty P.M., Jiang Z.-S., Liu H.-B., Degenhardt J., Mayba O., Gnad F., Liu J.-F., Pau G., Reeder J., Cao Y., Mukhyala K., Selvaraj S.K., Yu M.-M., Zynda G.J., Brauer M.J., Wu T.D., Gentleman R.C., Manning G., Yauch R.L., Bourgon R., Stokoe D., Modrusan Z., Neve R.M., de Sauvage F.J., Settleman J., Seshagiri S., Zhang Z.-M.
A comprehensive transcriptional portrait of human cancer cell lines.
Nat. Biotechnol. 33:306-312(2015)
PubMed=25846456; DOI=10.3892/ijo.2015.2951
Takenaka M., Saito M., Iwakawa R., Yanaihara N., Saito M., Kato M., Ichikawa H., Shibata T., Yokota J., Okamoto A., Kohno T.
Profiling of actionable gene alterations in ovarian cancer by targeted deep sequencing.
Int. J. Oncol. 46:2389-2398(2015)
PubMed=25877200; DOI=10.1038/nature14397
Yu M., Selvaraj S.K., Liang-Chu M.M.Y., Aghajani S., Busse M., Yuan J., Lee G., Peale F.V., Klijn C., Bourgon R., Kaminker J.S., Neve R.M.
A resource for cell line authentication, annotation and quality control.
Nature 520:307-311(2015)
PubMed=26169745; DOI=10.1186/s12967-015-0576-z; PMCID=PMC4499939
Halama A., Guerrouahen B.S., Pasquier J., Diboun I., Karoly E.D., Suhre K., Rafii A.
Metabolic signatures differentiate ovarian from colon cancer cell lines.
J. Transl. Med. 13:223.1-223.12(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5; PMCID=PMC4653878
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=26668597; DOI=10.3892/etm.2015.2836; PMCID=PMC4665172
Ruan Z.-Y., Liu J.-H., Kuang Y.-P.
Isolation and characterization of side population cells from the human ovarian cancer cell line SK-OV-3.
Exp. Ther. Med. 10:2071-2078(2015)
PubMed=26925792; DOI=10.4149/315_150930N510
Saczko J., Pilat J., Choromanska A., Rembialkowska N., Bar J.K., Deszcz I., Zalewski J., Kulbacka J.
The effectiveness of chemotherapy and electrochemotherapy on ovarian cell lines in vitro.
Neoplasma 63:450-455(2016)
PubMed=27235858; DOI=10.1016/j.ygyno.2016.05.028; PMCID=PMC4961516
Hernandez L., Kim M.K., Lyle L.T., Bunch K.P., House C.D., Ning F., Noonan A.M., Annunziata C.M.
Characterization of ovarian cancer cell lines as in vivo models for preclinical studies.
Gynecol. Oncol. 142:332-340(2016)
PubMed=27377824; DOI=10.1038/sdata.2016.52; PMCID=PMC4932877
Mestdagh P., Lefever S., Volders P.-J., Derveaux S., Hellemans J., Vandesompele J.
Long non-coding RNA expression profiling in the NCI60 cancer cell line panel using high-throughput RT-qPCR.
Sci. Data 3:160052-160052(2016)
PubMed=27397505; DOI=10.1016/j.cell.2016.06.017; PMCID=PMC4967469
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., Aben N., Goncalves E., Barthorpe S., Lightfoot H., Cokelaer T., Greninger P., van Dyk E., Chang H., de Silva H., Heyn H., Deng X.-M., Egan R.K., Liu Q.-S., Miroo T., Mitropoulos X., Richardson L., Wang J.-H., Zhang T.-H., Moran S., Sayols S., Soleimani M., Tamborero D., Lopez-Bigas N., Ross-Macdonald P., Esteller M., Gray N.S., Haber D.A., Stratton M.R., Benes C.H., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Garnett M.J.
A landscape of pharmacogenomic interactions in cancer.
Cell 166:740-754(2016)
PubMed=27807467; DOI=10.1186/s13100-016-0078-4; PMCID=PMC5087121
Zampella J.G., Rodic N., Yang W.R., Huang C.R.L., Welch J., Gnanakkan V.P., Cornish T.C., Boeke J.D., Burns K.H.
A map of mobile DNA insertions in the NCI-60 human cancer cell panel.
Mob. DNA 7:20.1-20.11(2016)
PubMed=28196595; DOI=10.1016/j.ccell.2017.01.005; PMCID=PMC5501076
Li J., Zhao W., Akbani R., Liu W.-B., Ju Z.-L., Ling S.-Y., Vellano C.P., Roebuck P., Yu Q.-H., Eterovic A.K., Byers L.A., Davies M.A., Deng W.-L., Gopal Y.N.V., Chen G., von Euw E.M., Slamon D.J., Conklin D., Heymach J.V., Gazdar A.F., Minna J.D., Myers J.N., Lu Y.-L., Mills G.B., Liang H.
Characterization of human cancer cell lines by reverse-phase protein arrays.
Cancer Cell 31:225-239(2017)
PubMed=28273451; DOI=10.1016/j.celrep.2017.02.028
Medrano M., Communal L., Brown K.R., Iwanicki M., Normand J., Paterson J., Sircoulomb F., Krzyzanowski P.M., Novak M., Doodnauth S.A., Suarez Saiz F.J., Cullis J., Al-awar R., Neel B.G., McPherson J., Drapkin R.I., Ailles L., Mes-Masson A.-M., Rottapel R.
Interrogation of functional cell-surface markers identifies CD151 dependency in high-grade serous ovarian cancer.
Cell Rep. 18:2343-2358(2017)
PubMed=30485824; DOI=10.1016/j.celrep.2018.10.096; PMCID=PMC6481945
Papp E., Hallberg D., Konecny G.E., Bruhm D.C., Adleff V., Noe M., Kagiampakis I., Palsgrove D., Conklin D., Kinose Y., White J.R., Press M.F., Drapkin R.I., Easwaran H., Baylin S.B., Slamon D.J., Velculescu V.E., Scharpf R.B.
Integrated genomic, epigenomic, and expression analyses of ovarian cancer cell lines.
Cell Rep. 25:2617-2633(2018)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747; PMCID=PMC6445675
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3; PMCID=PMC6697103
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. 3rd, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)"