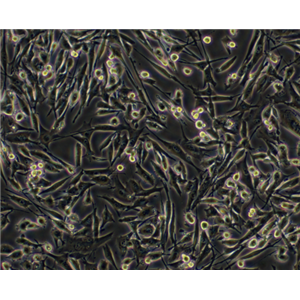
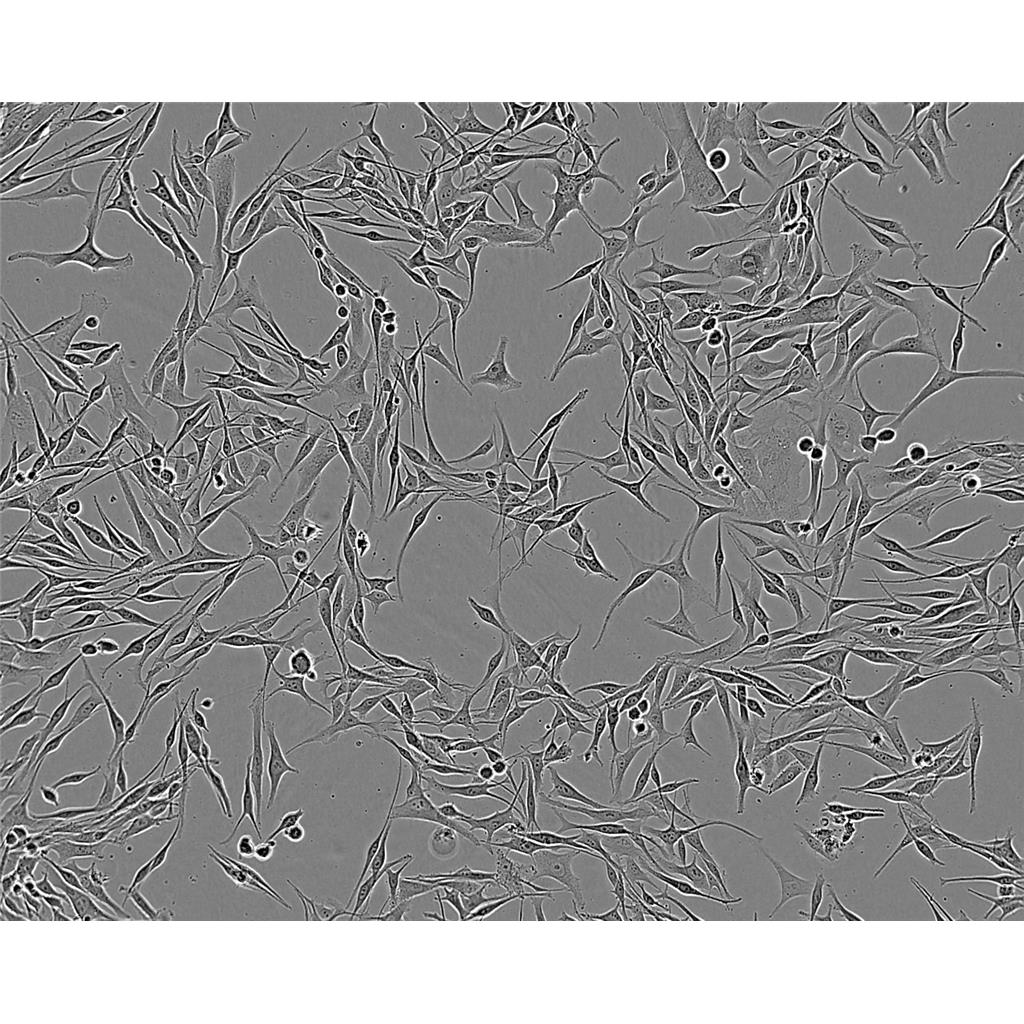
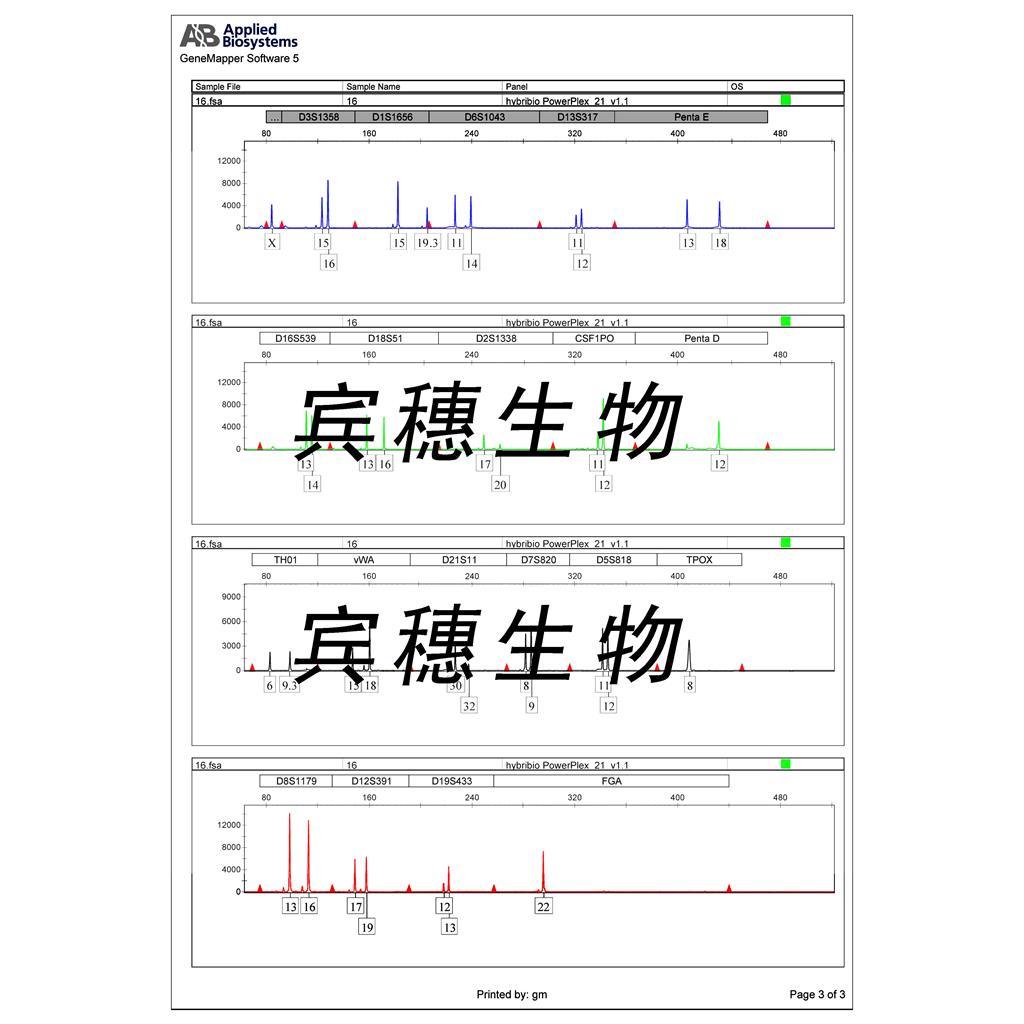
"DU 145人前列腺癌细胞代次低|培养基|送STR图谱
传代比例:1:2-1:4(首次传代建议1:2)
生长特性:贴壁生长
细胞系的应用:1)免疫组化研究2)RNA干扰研究3)药物作用研究4)慢病毒转染研究等其它应用。细胞系通常用于实验研究,如增殖、迁移、侵袭等。细胞系在多个领域的研究中被广泛应用,包括基础医学、临床试验、药物筛选和分子生物学研究。这些研究不仅在中国,也在日本、美国和欧洲等多个国家和地区进行。
换液周期:每周2-3次
GP2-293 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:AM38细胞、COLO320-DM细胞、NCI-H1944细胞
UACC 812 Cells;背景说明:该细胞是由Liebovitz A等于1986年从一名43岁的白人女性乳腺导管癌患者的乳腺切除肿瘤组织中分离建立的;手术前该病人曾接受过广泛的化疗。该细胞HER-2/neu癌基因序列有15倍的扩增;雌激素受体ER、孕激素受体PR和糖蛋白P阴性。;传代方法:1:3传代;5-7天1次。 ;生长特性:贴壁生长;形态特性:上皮样;相关产品有:RL-952细胞、VMM5细胞、Tu212细胞
SKNEP Cells;背景说明:超微结构有少许微绒毛,连接复合物,形态完整的高尔基体,内质网多为光滑型,脂滴,没有病毒棵粒。;传代方法:1:2传代。3天内可长满。;生长特性:半贴壁生长;形态特性:上皮细胞;相关产品有:U-138MG细胞、U14细胞、BIC1细胞
DU 145人前列腺癌细胞代次低|培养基|送STR图谱
背景信息:DU 145 是从一位有3年淋巴细胞白血病史的前列腺癌患者的脑部转移灶中建立的。该细胞系未检测到激素敏感性,性酶阳性,单个的细胞可在软琼脂中形成集落。对此细胞和原始肿瘤的亚显微结构分析可见微绒毛、微丝、细胞桥粒、线粒体、发达的高尔基体和异质溶酶体。该细胞不表达前列腺抗原。
┈订┈购(技术服务)┈热┈线:1┈3┈6┈4┈1┈9┈3┈0┈7┈9┈1【微信同号】┈Q┈Q:3┈1┈8┈0┈8┈0┈7┈3┈2┈4;
ATCC细胞库(American Type Culture Colection),该中心一直致力于细胞分类、鉴定和保藏工作。ATCC是全球最大的生物资源保藏中心,ATCC通过行业标准产品、服务和创新解决方案支持全球学术、政府、生物技术、制药、食品、农业和工业领域的科学进步。ATCC提供的服务和定制解决方案包括细胞和微生物培养、鉴定、生物衍生物的开发和生产、性能测试和生物资源保藏服务。美国国家标准协会(ANSI)认可了ATCC标准开发组织,并制定了标准协议,以确保生物材料的可靠性和可重复性。ATCC的使命是为了获取、鉴定、保存、开发、标准化和分发生物资源和生物信息,以提高和应用生物科学知识。
产品包装:复苏发货:T25培养瓶(一瓶)或冻存发货:1ml冻存管(两支)
来源说明:细胞主要来源ATCC、ECACC、DSMZ、RIKEN等细胞库
DU 145人前列腺癌细胞代次低|培养基|送STR图谱
物种来源:人源、鼠源等其它物种来源
Human Epidermoid carcinoma #2 Cells;背景说明:最初认为这个细胞源自喉上皮癌,但随后通过同功酶分析、HeLa标记染色体和DNA指纹分析发现,起源细胞已被HeLa污染。 角蛋白免疫过氧化物酶染色阳性。;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:HCC1569细胞、NBL-4细胞、NS20Y细胞
KPNRTBM1 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:成神经细胞;相关产品有:32D clone3细胞、H 9细胞、CP-70细胞
EOMA Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:P3/NSI/1-AG4-1细胞、OB2细胞、H-548细胞
LICCF Cells;背景说明:肝内胆管癌;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:Emory University-3细胞、DB细胞、EJ1细胞
┈订┈购(技术服务)┈热┈线:1┈3┈6┈4┈1┈9┈3┈0┈7┈9┈1【微信同号】┈Q┈Q:3┈1┈8┈0┈8┈0┈7┈3┈2┈4;
形态特性:上皮细胞样
贴壁细胞消化传代时通常采用两种方法:一、加入胰酶等细胞脱落后,再加培养基中止胰酶作用,离心传代;二、加入胰酶后,镜下观察待细胞始脱落时,弃胰酶,加培养分瓶。但前者太麻烦,而后者有可能对细胞施加胰酶选择,因为总是贴壁不牢的细胞先脱落,对肿瘤细胞来说,这部分细胞有可能是恶性程度较GAO的细胞亚群。一种简单的消化传代方法。加入PBS洗去血清或加入胰酶先中和血清的作用(30s),弃之,再加入适量胰酶作用10s-40s(根据细胞消化的难易程度),弃之,这样依赖残余的胰酶就可将细胞消化单细胞。对于较难消化的细胞,可以用2%利多卡因消化5-8分钟,然后再弃去,加培养基吹打也可以,对细胞的影响不大。不用PBS也不用Hanks洗,只要把旧培养吸的干净一点,直接加酶消化应该不会有什么问题。弃培养后,用0.04%的EDA冲洗一次,再用1/4v的0.04%的EDA室温孵育5min,弃取大部分EDA,加入与剩余EDA等量的胰酶(预热)总体积1/10v。消化到有细胞脱落。不过有人说EDA对细胞不HAO,有证据吗?培养的BASMC:倒掉旧培养加入少量胰酶冲一下,倒掉再加入0.125-0.25%胰酶约6-10滴或1ml(25ml bole)消化再加入适量新培养基中和,并分瓶这种方法简单、省事;效果很HAO并且不损失细胞!
OsACL Cells;背景说明:详见相关文献介绍;传代方法:1:5-1:10传代;每周换液2-3次。;生长特性:贴壁生长;形态特性:成纤维细胞;相关产品有:Instituto Biologico-Rim Suino-2细胞、LU65细胞、SUM-159-PT细胞
SJ-Rh 30 Cells;背景说明:肺泡横纹肌肉瘤;骨髓转移;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:LLC PK1细胞、H716细胞、RKN细胞
EB1 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:悬浮生长 ;形态特性:淋巴母细胞样;相关产品有:Colo320细胞、CCFSTTG1细胞、J774A1细胞
DOK Cells;背景说明:口腔异常增生;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:GM12878细胞、Nthy ori 3.1细胞、NB-19细胞
NCI-H2342 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:6传代 ;生长特性:贴壁生长;形态特性:上皮样;相关产品有:174xCEM.T2细胞、NCIH889细胞、hEEC细胞
COLO-357 Cells;背景说明:胰腺癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:SNU449细胞、S91细胞、MM1.S细胞
NCM356 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:MDCK II细胞、REC1细胞、AK细胞
Hs 274 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代,每周换液2-3次。;生长特性:贴壁生长;形态特性:成纤维细胞;相关产品有:CFPAC-1细胞、TK-10细胞、AML 12细胞
Duke University 4475 Cells;背景说明:详见相关文献介绍;传代方法:每周换液2—3次;生长特性:悬浮,多细胞聚集;形态特性:上皮细胞样;相关产品有:SUM-190PT细胞、EM3细胞、CMT64细胞
KYSE 270 Cells;背景说明:详见相关文献介绍;传代方法:1:5传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:HOP92细胞、Hs739T细胞、G-361细胞
NCI-H841 Cells;背景说明:详见相关文献介绍;传代方法:1:3—1:5传代,;生长特性:混合型生长;形态特性:详见产品说明书;相关产品有:BNL.1ME A.7R.1细胞、NOZAWA细胞、Colo16细胞
NCI-H1668 Cells;背景说明:小细胞肺癌;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:NPA细胞、Michigan Cancer Foundation-7细胞、HOS TE85细胞
Hs 863.T Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代;每周换液2-3次。;生长特性:贴壁生长;形态特性:成纤维细胞;相关产品有:CHO-Lec1细胞、A431细胞、WEHI-231细胞
UMUC1 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:HLE细胞、HFL-1细胞、H-2029细胞
BEAS2B Cells;背景说明:从一位非癌个体的正常人支气管上皮病理切片分离出上皮细胞。这些细胞用腺病毒12-SV40病毒杂交病毒感染并克隆。DEAS-2B细胞保留了对血清反应进行鳞关分化的能力,并有用于筛选诱导或影响分化及致癌的化学或生物制剂。细胞角蛋白及SV40抗原染色阳性。;传代方法:消化3-5分钟。1:2。3天内可长满;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:JOSK-M细胞、MCA 38细胞、WM-239-A细胞
Jijoye Cells;背景说明:详见相关文献介绍;传代方法:每周2-3次。;生长特性:悬浮生长;形态特性:淋巴母细胞;相关产品有:D283细胞、SRA 01/04细胞、Mo 59J细胞
HS-5 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:上皮细胞样;相关产品有:HEK293-F细胞、HEK293细胞、TN5B14细胞
Abcam A-549 KAT2A KO Cells(提供STR鉴定图谱)
Abcam U-87MG KCND2 KO Cells(提供STR鉴定图谱)
BayGenomics ES cell line CSG100 Cells(提供STR鉴定图谱)
BayGenomics ES cell line RRT487 Cells(提供STR鉴定图谱)
BayGenomics ES cell line YTC481 Cells(提供STR鉴定图谱)
CHO IM4 Cells(提供STR鉴定图谱)
DA02346 Cells(提供STR鉴定图谱)
F9-41 Cells(提供STR鉴定图谱)
GM07748 Cells(提供STR鉴定图谱)
HCC0078 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:HT144细胞、OVCAR-4细胞、HRCEC细胞
DU 145人前列腺癌细胞代次低|培养基|送STR图谱
NCI-H647 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代;每周换液2次。;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:HCCLM6细胞、SPC-A-1细胞、Intestinal Epithelioid Cell line No. 6细胞
SNGM Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:多边形;相关产品有:H2009细胞、RT4-D6-P2T细胞、293-FT细胞
Central Adrenergic TH-expressing a Cells;背景说明:神经;SV40转化;C57BL/6 x DBA/2;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:MPC-11细胞、H-128细胞、LAPC4细胞
H2196 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代;每周换液2次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:LA-N-6(OAN)细胞、MC/CAR细胞、IGR-OV1细胞
NP69SV40T Cells;背景说明:鼻咽;上皮细胞;SV40转化;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:MC3T3E1细胞、HOEC细胞、MGHU1细胞
B16-A Cells(提供STR鉴定图谱)
U-2932 Cells;背景说明:弥漫大B淋巴瘤;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:Hs 839.T细胞、P3HR1细胞、NRK52E细胞
T98G Cells;背景说明:详见相关文献介绍;传代方法:按1:3传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:RS4:11细胞、130T细胞、NSI/1-Ag4-1细胞
Molm14 Cells;背景说明:急性髓系白血病;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:CAL78细胞、HCE-2 [50.B1]细胞、HS 445T细胞
BN-CL2 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:SW-1271细胞、OCI/AML-4细胞、Calu-3细胞
SNK6 Cells;背景说明:NK/T细胞淋巴瘤;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:BJA-B1细胞、H69C细胞、CCC-HPF-1细胞
MRASMC Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Tohoku Hospital Pediatrics-1细胞、EC9706细胞、CCD-18Co细胞
HT 1080.T Cells;背景说明:该细胞源自一名35岁患有纤维肉瘤的白人男性的结缔组织;ras+。;传代方法:1:4-1:8传代;2-3天换液1次。;生长特性:贴壁生长;形态特性:上皮样;相关产品有:SK-Hep1细胞、HANK细胞、HFL细胞
IPLB-SF 21AE Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:SKO3细胞、Intestinal Porcine Epithelial Cell line-J2细胞、HCC-2108细胞
GM10123 Cells(提供STR鉴定图谱)
HAP1 C16orf91 (-) 1 Cells(提供STR鉴定图谱)
EJ 138 Cells;背景说明:膀胱癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:IEC18细胞、MV(4;11)细胞、MSCs(mUCMSCs)细胞
TW-039 Cells;背景说明:鼻咽癌;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:RT-4细胞、102PT细胞、GalK 1细胞
NCI-H522 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代;每周换液2-3次。;生长特性:贴壁生长;形态特性:上皮样;相关产品有:SJCRH30细胞、RIN-m 14B细胞、H2170细胞
OCI-Ly3 Cells;背景说明:弥漫大B淋巴瘤;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:C8166细胞、WERI-Rb-1细胞、PLA801D细胞
RPMI-8226S Cells;背景说明:来源于一位61岁的男性浆细胞瘤患者;可产生免疫球蛋白轻链,未检测到重链。;传代方法:按1:2传代,5-6小时可以看到细胞分裂;生长特性:悬浮生长;形态特性:淋巴母细胞样;相关产品有:RCSMC细胞、C32 mel细胞、HT115细胞
NTERA-2 cl.D1 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:GM02219细胞、HMC3细胞、B16/F1细胞
KMST6 Cells;背景说明:胚成纤维细胞;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:MH7A细胞、SUM102PT细胞、NB19-RIKEN细胞
293EBNA Cells;背景说明:详见相关文献介绍;传代方法:1:4-1:10传代;每周2次。;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:GM-3573细胞、A375-MEL细胞、Hs-281-T细胞
HG03549 Cells(提供STR鉴定图谱)
IDG-HEK293T-GRID1-isoform-1-V5-OE Cells(提供STR鉴定图谱)
LPC-Aa98-19 Cells(提供STR鉴定图谱)
NCI-HPN-F1K Colon Cells(提供STR鉴定图谱)
PathHunter U2OS OPRM1 beta-arrestin Cells(提供STR鉴定图谱)
Ubigene HCT 116 SMAD6 KO Cells(提供STR鉴定图谱)
WAe009-A-52 Cells(提供STR鉴定图谱)
HG01089 Cells(提供STR鉴定图谱)
J 111 Cells;背景说明:单核细胞白血病;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:NCI-H2030细胞、MRMT-1细胞、L929细胞
CAL 12T Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:HOS-TE85细胞、SW-1990细胞、MES-SA/Dx-5细胞
AN3 CA Cells;背景说明:AN3CA细胞建系于1964年。它衍生于子宫内膜癌患者淋巴结转移组织,具有癌细胞的基本特性,能在体外长期传代培养,接种实验动物产生明显肿瘤。但细胞的生物学特性及超微结构尚未深入研究,仅发现该细胞系促黑激素合成为阴性。细胞常用于人子宫内膜癌细胞生物学及其相关特性研究。;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:BV-2细胞、Ishikawa细胞、COLO 679细胞
MDCC-MSB-1 Cells;背景说明:淋巴瘤;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:C-33-A细胞、RDES-1细胞、NCI-H1623细胞
Chang Cells Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Z138细胞、G-361细胞、KYSE450细胞
DMS 79 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代;每周换液2-3次。;生长特性:悬浮生长;形态特性:详见产品说明书;相关产品有:SH-SY5Y Parental细胞、Cates-1B细胞、HN6细胞
HeLa 229 Cells;背景说明:宫颈癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:NTERA-2cl.D1细胞、LS 180细胞、FRTL5细胞
MC3T3E1 Cells;背景说明:该细胞有多个亚克隆,可以作为体外研究成骨细胞分化的良好模型,尤其是ECM信号通路的作用。;传代方法:1:2传代;生长特性:贴壁生长;形态特性:成纤维细胞样;相关产品有:Panc10.05细胞、RH7777细胞、UT7细胞
MC-116 Cells;背景说明:详见相关文献介绍;传代方法:每周换液2-3次。;生长特性:悬浮生长 ;形态特性:淋巴母细胞样;相关产品有:SBC3细胞、NPA87-1细胞、H-510细胞
NCI-H1522 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:H441细胞、GM03671细胞、SK-HEP-1细胞
DMS79 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代;每周换液2-3次。;生长特性:悬浮生长;形态特性:详见产品说明书;相关产品有:RBL-1细胞、SK MES 1细胞、BGC-823细胞
DMS-79 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代;每周换液2-3次。;生长特性:悬浮生长;形态特性:详见产品说明书;相关产品有:U031细胞、AM38细胞、UMUC1细胞
hTERTHME1 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:SK-MEL-3细胞、PaCa2细胞、LY-R细胞
BT 549 Cells;背景说明:该细胞1978年由W.G.Coutinho和E.Y.Lasfargues建系,源自一位72岁患有乳腺导管癌的白人女性,来源组织包括乳头及浸润导管。该细胞形态包括上皮样细胞及多核巨细胞,可分泌一种粘性物质。;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:DU4475细胞、HS-729细胞、Mouse podocyte细胞
HEK293-EBNA Cells;背景说明:详见相关文献介绍;传代方法:1:4-1:10传代;每周2次。;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:Panc 2.03细胞、Hs746T细胞、U-937细胞
SBG5 Cells(提供STR鉴定图谱)
HEB Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:KMS26细胞、H-847细胞、EB1细胞
SKOV3 Cells;背景说明:SK-OV-3由G.Trempe和L.J.Old在1973年从卵巢肿瘤病人的腹水分离得到。 此细胞对肿瘤坏死因子和几种细胞毒性药物包括白喉毒素、顺铂和阿霉素均耐受。 在裸鼠中致瘤,且形成与卵巢原位癌一致的中度分化的腺癌。;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:上皮细胞样;相关产品有:MDA 1386细胞、3T3-L1细胞、SW-1088细胞
HO-8910 PM Cells;背景说明:高转移卵巢癌 Cells;传代方法:消化3-5分钟。1:2。3天内可长满。;生长特性:贴壁生长;形态特性:上皮样;相关产品有:CL-40细胞、H-2227细胞、NCI-H345细胞
NCI-SNU-1040 Cells;背景说明:结肠癌;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:DHL-2细胞、NIE 115细胞、H-1417细胞
NCI-H1836 Cells;背景说明:详见相关文献介绍;传代方法:每周换液2-3次。;生长特性:悬浮生长;形态特性:详见产品说明书;相关产品有:GRSL细胞、WM 266-4细胞、Jurkat 77细胞
PG-LH7 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:RSMC细胞、T/GHA-VSMC细胞、CAL 78细胞
DU 145人前列腺癌细胞代次低|培养基|送STR图谱
MLTC-1 Cells;背景说明:睾丸间质瘤;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:MV3细胞、HemECs细胞、SK N SH细胞
H-2195 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:H-226细胞、Hs683细胞、D324 Med细胞
KGN Cells;背景说明:该细胞株是颗粒细胞瘤。细胞在加入人体绒膜促性腺激素后可能产生孕酮。细胞生长缓慢。;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:上皮细胞样;相关产品有:SO-RB50细胞、HS688AT细胞、SKCO1细胞
SW-1783 Cells;背景说明:间变性星形细胞瘤;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:FRhK4细胞、Jeko1细胞、G 401细胞
SUM-102 Cells;背景说明:乳腺癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:H-1651细胞、IPAM-WT细胞、ACC-2细胞
RIN-m5F Cells;背景说明:胰岛β细胞瘤;雄性;NEDH;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:NCIN87细胞、MCF 7细胞、TE11细胞
HPDE Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:MDCC MSB1细胞、CA922细胞、CPA 47细胞
PK 15 Cells;背景说明:PK-15细胞建系于1955(Stice,E)。是PK-1a细胞的克隆系。该细胞系可用于多种病毒的增值及特性研究。另外,电镜观察发现,PK-15细胞内有C-型病毒颗粒存在,是研究C-型病毒的材料。;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:上皮细胞样;相关产品有:HF-91细胞、A172细胞、NOMO-1细胞
BayGenomics ES cell line KST216 Cells(提供STR鉴定图谱)
BayGenomics ES cell line XB777 Cells(提供STR鉴定图谱)
Col-10 Cells(提供STR鉴定图谱)
MANFU2-10F9 Cells(提供STR鉴定图谱)
S2G8 Cells(提供STR鉴定图谱)
YTH 71.3 Cells(提供STR鉴定图谱)
" "PubMed=7017212; DOI=10.1093/jnci/66.6.1003
Pollack M.S., Heagney S.D., Livingston P.O., Fogh J.
HLA-A, B, C and DR alloantigen expression on forty-six cultured human tumor cell lines.
J. Natl. Cancer Inst. 66:1003-1012(1981)
PubMed=3518877; DOI=10.3109/07357908609038260
Fogh J.
Human tumor lines for cancer research.
Cancer Invest. 4:157-184(1986)
PubMed=3335022
Alley M.C., Scudiero D.A., Monks A., Hursey M.L., Czerwinski M.J., Fine D.L., Abbott B.J., Mayo J.G., Shoemaker R.H., Boyd M.R.
Feasibility of drug screening with panels of human tumor cell lines using a microculture tetrazolium assay.
Cancer Res. 48:589-601(1988)
PubMed=2744886; DOI=10.1002/ijc.2910440128
Hartley-Asp B., Billstrom A., Kruse E.
Identification by C-banding of two human prostate tumour cell lines, 1013L and DU 145.
Int. J. Cancer 44:161-164(1989)
PubMed=1873816
Isaacs W.B., Carter B.S., Ewing C.M.
Wild-type p53 suppresses growth of human prostate cancer cells containing mutant p53 alleles.
Cancer Res. 51:4716-4720(1991)
PubMed=1574572; DOI=10.2307/3578273
Dunphy E.J., Beckett M.A., Thompson L.H., Weichselbaum R.R.
Expression of the polymorphic human DNA repair gene XRCC1 does not correlate with radiosensitivity in the cells of human head and neck tumor cell lines.
Radiat. Res. 130:166-170(1992)
PubMed=8104329; DOI=10.1002/pros.2990230206
Carroll A.G., Voeller H.J., Sugars L., Gelmann E.P.
p53 oncogene mutations in three human prostate cancer cell lines.
Prostate 23:123-134(1993)
PubMed=8510267; DOI=10.1016/S0022-5347(17)35458-7
Effert P.J., McCoy R.H., Walther P.J., Liu E.T.-B.
p53 gene alterations in human prostate carcinoma.
J. Urol. 150:257-261(1993)
PubMed=9018337; DOI=10.1002/(SICI)1097-0045(19970101)30:1<58::AID-PROS9>3.0.CO;2-H
Webber M.M., Bello D., Quader S.T.A.
Immortalized and tumorigenic adult human prostatic epithelial cell lines: characteristics and applications Part 2. Tumorigenic cell lines.
Prostate 30:58-64(1997)
PubMed=9214606; DOI=10.1093/carcin/18.6.1225
Webber M.M., Bello D., Kleinman H.K., Hoffman M.P.
Acinar differentiation by non-malignant immortalized human prostatic epithelial cells and its loss by malignant cells.
Carcinogenesis 18:1225-1231(1997)
PubMed=9460501; DOI=10.1016/S0165-4608(97)00060-5
Nupponen N.N., Hyytinen E.-R., Kallioniemi A.H., Visakorpi T.
Genetic alterations in prostate cancer cell lines detected by comparative genomic hybridization.
Cancer Genet. Cytogenet. 101:53-57(1998)
PubMed=10702678; DOI=10.1159/000015432
Pan Y., Kytola S., Farnebo F., Wang N., Lui W.-O., Nupponen N.N., Isola J.J., Visakorpi T., Bergerheim U.S.R., Larsson C.
Characterization of chromosomal abnormalities in prostate cancer cell lines by spectral karyotyping.
Cytogenet. Cell Genet. 87:225-232(1999)
PubMed=10700174; DOI=10.1038/73432
Ross D.T., Scherf U., Eisen M.B., Perou C.M., Rees C., Spellman P.T., Iyer V.R., Jeffrey S.S., van de Rijn M., Waltham M.C., Pergamenschikov A., Lee J.C.F., Lashkari D., Shalon D., Myers T.G., Weinstein J.N., Botstein D., Brown P.O.
Systematic variation in gene expression patterns in human cancer cell lines.
Nat. Genet. 24:227-235(2000)
PubMed=10972993; DOI=10.1002/1098-2744(200008)28:4<236::AID-MC6>3.0.CO;2-H
Rauh-Adelmann C., Lau K.-M., Sabeti N., Long J.P., Mok S.C., Ho S.-M.
Altered expression of BRCA1, BRCA2, and a newly identified BRCA2 exon 12 deletion variant in malignant human ovarian, prostate, and breast cancer cell lines.
Mol. Carcinog. 28:236-246(2000)
PubMed=11135431; DOI=10.1002/1098-2264(2000)9999:9999<::AID-GCC1076>3.0.CO;2-E
Aurich-Costa J., Vannier A., Gregoire E., Nowak F., Cherif D.
IPM-FISH, a new M-FISH approach using IRS-PCR painting probes: application to the analysis of seven human prostate cell lines.
Genes Chromosomes Cancer 30:143-160(2001)
PubMed=11172901; DOI=10.1016/S0165-4608(00)00339-3
Strefford J.C., Lillington D.M., Young B.D., Oliver R.T.D.
The use of multicolor fluorescence technologies in the characterization of prostate carcinoma cell lines: a comparison of multiplex fluorescence in situ hybridization and spectral karyotyping data.
Cancer Genet. Cytogenet. 124:112-121(2001)
PubMed=11280753
Urasaki Y., Laco G.S., Pourquier P., Takebayashi Y., Kohlhagen G., Gioffre C., Zhang H.-L., Chatterjee D., Pantazis P., Pommier Y.
Characterization of a novel topoisomerase I mutation from a camptothecin-resistant human prostate cancer cell line.
Cancer Res. 61:1964-1969(2001)
PubMed=11304728; DOI=10.1002/pros.1045
van Bokhoven A., Varella-Garcia M., Korch C.T., Hessels D., Miller G.J.
Widely used prostate carcinoma cell lines share common origins.
Prostate 47:36-51(2001)
PubMed=11414198; DOI=10.1007/s004320000207
Lahm H., Andre S., Hoeflich A., Fischer J.R., Sordat B., Kaltner H., Wolf E., Gabius H.-J.
Comprehensive galectin fingerprinting in a panel of 61 human tumor cell lines by RT-PCR and its implications for diagnostic and therapeutic procedures.
J. Cancer Res. Clin. Oncol. 127:375-386(2001)
PubMed=11416159; DOI=10.1073/pnas.121616198; PMCID=PMC35459
Masters J.R.W., Thomson J.A., Daly-Burns B., Reid Y.A., Dirks W.G., Packer P., Toji L.H., Ohno T., Tanabe H., Arlett C.F., Kelland L.R., Harrison M., Virmani A.K., Ward T.H., Ayres K.L., Debenham P.G.
Short tandem repeat profiling provides an international reference standard for human cell lines.
Proc. Natl. Acad. Sci. U.S.A. 98:8012-8017(2001)
PubMed=12606952; DOI=10.1038/sj.onc.1206247
Clark J., Edwards S., Feber A., Flohr P., John M., Giddings I., Crossland S., Stratton M.R., Wooster R., Campbell C., Cooper C.S.
Genome-wide screening for complete genetic loss in prostate cancer by comparative hybridization onto cDNA microarrays.
Oncogene 22:1247-1252(2003)
PubMed=12615715
Reinhold W.C., Kouros-Mehr H., Kohn K.W., Maunakea A.K., Lababidi S., Roschke A.V., Stover K., Alexander J., Pantazis P., Miller L., Liu E.T.-B., Kirsch I.R., Urasaki Y., Pommier Y., Weinstein J.N.
Apoptotic susceptibility of cancer cells selected for camptothecin resistance: gene expression profiling, functional analysis, and molecular interaction mapping.
Cancer Res. 63:1000-1011(2003)
PubMed=12725112; DOI=10.1385/1-59259-372-0:21
Russell P.J., Kingsley E.A.
Human prostate cancer cell lines.
Methods Mol. Med. 81:21-39(2003)
PubMed=14518029; DOI=10.1002/pros.10290
van Bokhoven A., Varella-Garcia M., Korch C.T., Johannes W.U., Smith E.E., Miller H.L., Nordeen S.K., Miller G.J., Lucia M.S.
Molecular characterization of human prostate carcinoma cell lines.
Prostate 57:205-225(2003)
CLPUB00698
van Bokhoven A.
Models for prostate cancer. Molecular characterization and critical appraisal of human prostate carcinoma cell lines.
Thesis PhD (2004); Katholieke Universiteit Nijmegen; Nijmegen; Netherlands
PubMed=15486987; DOI=10.1002/pros.20158
Zhao H.-J., Kim Y., Wang P., Lapointe J., Tibshirani R., Pollack J.R., Brooks J.D.
Genome-wide characterization of gene expression variations and DNA copy number changes in prostate cancer cell lines.
Prostate 63:187-197(2005)
PubMed=15748285; DOI=10.1186/1479-5876-3-11; PMCID=PMC555742
Adams S., Robbins F.-M., Chen D., Wagage D., Holbeck S.L., Morse H.C. 3rd, Stroncek D., Marincola F.M.
HLA class I and II genotype of the NCI-60 cell lines.
J. Transl. Med. 3:11.1-11.8(2005)
PubMed=17088437; DOI=10.1158/1535-7163.MCT-06-0433; PMCID=PMC2705832
Ikediobi O.N., Davies H.R., Bignell G.R., Edkins S., Stevens C., O'Meara S., Santarius T., Avis T., Barthorpe S., Brackenbury L., Buck G., Butler A.P., Clements J., Cole J., Dicks E., Forbes S., Gray K., Halliday K., Harrison R., Hills K., Hinton J., Hunter C., Jenkinson A., Jones D., Kosmidou V., Lugg R., Menzies A., Miroo T., Parker A., Perry J., Raine K.M., Richardson D., Shepherd R., Small A., Smith R., Solomon H., Stephens P.J., Teague J.W., Tofts C., Varian J., Webb T., West S., Widaa S., Yates A., Reinhold W.C., Weinstein J.N., Stratton M.R., Futreal P.A., Wooster R.
Mutation analysis of 24 known cancer genes in the NCI-60 cell line set.
Mol. Cancer Ther. 5:2606-2612(2006)
PubMed=17254797; DOI=10.1016/j.biologicals.2006.10.001
Azari S., Ahmadi N., Jeddi-Tehrani M., Shokri F.
Profiling and authentication of human cell lines using short tandem repeat (STR) loci: report from the National Cell Bank of Iran.
Biologicals 35:195-202(2007)
PubMed=17440963; DOI=10.1002/pros.20581
Takeda M., Mizokami A., Mamiya K., Li Y.-Q., Zhang J., Keller E.T., Namiki M.
The establishment of two paclitaxel-resistant prostate cancer cell lines and the mechanisms of paclitaxel resistance with two cell lines.
Prostate 67:955-967(2007)
PubMed=19372543; DOI=10.1158/1535-7163.MCT-08-0921; PMCID=PMC4020356
Lorenzi P.L., Reinhold W.C., Varma S., Hutchinson A.A., Pommier Y., Chanock S.J., Weinstein J.N.
DNA fingerprinting of the NCI-60 cell line panel.
Mol. Cancer Ther. 8:713-724(2009)
PubMed=20164919; DOI=10.1038/nature08768; PMCID=PMC3145113
Bignell G.R., Greenman C.D., Davies H.R., Butler A.P., Edkins S., Andrews J.M., Buck G., Chen L., Beare D., Latimer C., Widaa S., Hinton J., Fahey C., Fu B.-Y., Swamy S., Dalgliesh G.L., Teh B.T., Deloukas P., Yang F.-T., Campbell P.J., Futreal P.A., Stratton M.R.
Signatures of mutation and selection in the cancer genome.
Nature 463:893-898(2010)
PubMed=22068913; DOI=10.1073/pnas.1111840108; PMCID=PMC3219108
Gillet J.-P., Calcagno A.M., Varma S., Marino M., Green L.J., Vora M.I., Patel C., Orina J.N., Eliseeva T.A., Singal V., Padmanabhan R., Davidson B., Ganapathi R., Sood A.K., Rueda B.R., Ambudkar S.V., Gottesman M.M.
Redefining the relevance of established cancer cell lines to the study of mechanisms of clinical anti-cancer drug resistance.
Proc. Natl. Acad. Sci. U.S.A. 108:18708-18713(2011)
PubMed=22275356; DOI=10.1074/jbc.M111.302547; PMCID=PMC3322861
Lynch T.P., Ferrer C.M., Jackson S.R., Shahriari K.S., Vosseller K., Reginato M.J.
Critical role of O-linked beta-N-acetylglucosamine transferase in prostate cancer invasion, angiogenesis, and metastasis.
J. Biol. Chem. 287:11070-11081(2012)
PubMed=22336246; DOI=10.1016/j.bmc.2012.01.017
Kong D.-X., Yamori T.
JFCR39, a panel of 39 human cancer cell lines, and its application in the discovery and development of anticancer drugs.
Bioorg. Med. Chem. 20:1947-1951(2012)
PubMed=22347499; DOI=10.1371/journal.pone.0031628; PMCID=PMC3276511
Ruan X.-Y., Kocher J.-P.A., Pommier Y., Liu H.-F., Reinhold W.C.
Mass homozygotes accumulation in the NCI-60 cancer cell lines as compared to HapMap trios, and relation to fragile site location.
PLoS ONE 7:E31628-E31628(2012)
PubMed=22384151; DOI=10.1371/journal.pone.0032096; PMCID=PMC3285665
Lee J.-S., Kim Y.K., Kim H.J., Hajar S., Tan Y.L., Kang N.-Y., Ng S.H., Yoon C.N., Chang Y.-T.
Identification of cancer cell-line origins using fluorescence image-based phenomic screening.
PLoS ONE 7:E32096-E32096(2012)
PubMed=22460905; DOI=10.1038/nature11003; PMCID=PMC3320027
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=22628656; DOI=10.1126/science.1218595; PMCID=PMC3526189
Jain M., Nilsson R., Sharma S., Madhusudhan N., Kitami T., Souza A.L., Kafri R., Kirschner M.W., Clish C.B., Mootha V.K.
Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation.
Science 336:1040-1044(2012)
PubMed=23671654; DOI=10.1371/journal.pone.0063056; PMCID=PMC3646030
Lu Y.-H., Soong T.D., Elemento O.
A novel approach for characterizing microsatellite instability in cancer cells.
PLoS ONE 8:E63056-E63056(2013)
PubMed=23856246; DOI=10.1158/0008-5472.CAN-12-3342; PMCID=PMC4893961
Abaan O.D., Polley E.C., Davis S.R., Zhu Y.-L.J., Bilke S., Walker R.L., Pineda M.A., Gindin Y., Jiang Y., Reinhold W.C., Holbeck S.L., Simon R.M., Doroshow J.H., Pommier Y., Meltzer P.S.
The exomes of the NCI-60 panel: a genomic resource for cancer biology and systems pharmacology.
Cancer Res. 73:4372-4382(2013)
PubMed=23933261; DOI=10.1016/j.celrep.2013.07.018
Moghaddas Gholami A., Hahne H., Wu Z.-X., Auer F.J., Meng C., Wilhelm M., Kuster B.
Global proteome analysis of the NCI-60 cell line panel.
Cell Rep. 4:609-620(2013)
PubMed=23957452; DOI=10.3109/01913123.2013.814739
Gilloteaux J., Eze N., Jamison J.M., McGuire K., Summers J.L.
A rare, human prostate oncocyte cell originates from the prostatic carcinoma (DU145) cell line.
Ultrastruct. Pathol. 37:440-448(2013)
PubMed=24279929; DOI=10.1186/2049-3002-1-20; PMCID=PMC4178206
Dolfi S.C., Chan L.L.-Y., Qiu J., Tedeschi P.M., Bertino J.R., Hirshfield K.M., Oltvai Z.N., Vazquez A.
The metabolic demands of cancer cells are coupled to their size and protein synthesis rates.
Cancer Metab. 1:20.1-20.13(2013)
PubMed=24670534; DOI=10.1371/journal.pone.0092047; PMCID=PMC3966786
Varma S., Pommier Y., Sunshine M., Weinstein J.N., Reinhold W.C.
High resolution copy number variation data in the NCI-60 cancer cell lines from whole genome microarrays accessible through CellMiner.
PLoS ONE 9:E92047-E92047(2014)
PubMed=25485619; DOI=10.1038/nbt.3080
Klijn C., Durinck S., Stawiski E.W., Haverty P.M., Jiang Z.-S., Liu H.-B., Degenhardt J., Mayba O., Gnad F., Liu J.-F., Pau G., Reeder J., Cao Y., Mukhyala K., Selvaraj S.K., Yu M.-M., Zynda G.J., Brauer M.J., Wu T.D., Gentleman R.C., Manning G., Yauch R.L., Bourgon R., Stokoe D., Modrusan Z., Neve R.M., de Sauvage F.J., Settleman J., Seshagiri S., Zhang Z.-M.
A comprehensive transcriptional portrait of human cancer cell lines.
Nat. Biotechnol. 33:306-312(2015)
PubMed=25877200; DOI=10.1038/nature14397
Yu M., Selvaraj S.K., Liang-Chu M.M.Y., Aghajani S., Busse M., Yuan J., Lee G., Peale F.V., Klijn C., Bourgon R., Kaminker J.S., Neve R.M.
A resource for cell line authentication, annotation and quality control.
Nature 520:307-311(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5; PMCID=PMC4653878
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=26972028; DOI=10.1016/j.jprot.2016.03.008
Masuishi Y., Kimura Y., Arakawa N., Hirano H.
Identification of glycosylphosphatidylinositol-anchored proteins and omega-sites using TiO2-based affinity purification followed by hydrogen fluoride treatment.
J. Proteomics 139:77-83(2016)
PubMed=27141528; DOI=10.1016/j.dib.2016.04.001; PMCID=PMC4838930
Masuishi Y., Kimura Y., Arakawa N., Hirano H.
Data for identification of GPI-anchored peptides and omega-sites in cancer cell lines.
Data Brief 7:1302-1305(2016)
PubMed=27377824; DOI=10.1038/sdata.2016.52; PMCID=PMC4932877
Mestdagh P., Lefever S., Volders P.-J., Derveaux S., Hellemans J., Vandesompele J.
Long non-coding RNA expression profiling in the NCI60 cancer cell line panel using high-throughput RT-qPCR.
Sci. Data 3:160052-160052(2016)
PubMed=27397505; DOI=10.1016/j.cell.2016.06.017; PMCID=PMC4967469
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., Aben N., Goncalves E., Barthorpe S., Lightfoot H., Cokelaer T., Greninger P., van Dyk E., Chang H., de Silva H., Heyn H., Deng X.-M., Egan R.K., Liu Q.-S., Miroo T., Mitropoulos X., Richardson L., Wang J.-H., Zhang T.-H., Moran S., Sayols S., Soleimani M., Tamborero D., Lopez-Bigas N., Ross-Macdonald P., Esteller M., Gray N.S., Haber D.A., Stratton M.R., Benes C.H., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Garnett M.J.
A landscape of pharmacogenomic interactions in cancer.
Cell 166:740-754(2016)
PubMed=27807467; DOI=10.1186/s13100-016-0078-4; PMCID=PMC5087121
Zampella J.G., Rodic N., Yang W.R., Huang C.R.L., Welch J., Gnanakkan V.P., Cornish T.C., Boeke J.D., Burns K.H.
A map of mobile DNA insertions in the NCI-60 human cancer cell panel.
Mob. DNA 7:20.1-20.11(2016)
PubMed=28196595; DOI=10.1016/j.ccell.2017.01.005; PMCID=PMC5501076
Li J., Zhao W., Akbani R., Liu W.-B., Ju Z.-L., Ling S.-Y., Vellano C.P., Roebuck P., Yu Q.-H., Eterovic A.K., Byers L.A., Davies M.A., Deng W.-L., Gopal Y.N.V., Chen G., von Euw E.M., Slamon D.J., Conklin D., Heymach J.V., Gazdar A.F., Minna J.D., Myers J.N., Lu Y.-L., Mills G.B., Liang H.
Characterization of human cancer cell lines by reverse-phase protein arrays.
Cancer Cell 31:225-239(2017)
PubMed=30244336; DOI=10.1007/s00345-018-2501-6
Samli H., Samli M., Vatansever B., Ardicli S., Aztopal N., Dincel D., Sahin A., Balci F.
Paclitaxel resistance and the role of miRNAs in prostate cancer cell lines.
World J. Urol. 37:1117-1126(2019)
PubMed=30629668; DOI=10.1371/journal.pone.0210404; PMCID=PMC6328144
Uphoff C.C., Pommerenke C., Denkmann S.A., Drexler H.G.
Screening human cell lines for viral infections applying RNA-Seq data analysis.
PLoS ONE 14:E0210404-E0210404(2019)
PubMed=30787054; DOI=10.1158/1055-9965.EPI-18-1132; PMCID=PMC6548687
Hooker S.E. Jr., Woods-Burnham L., Bathina M., Lloyd S., Gorjala P., Mitra R., Nonn L., Kimbro K.S., Kittles R.A.
Genetic ancestry analysis reveals misclassification of commonly used cancer cell lines.
Cancer Epidemiol. Biomarkers Prev. 28:1003-1009(2019)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747; PMCID=PMC6445675
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=30971826; DOI=10.1038/s41586-019-1103-9
Behan F.M., Iorio F., Picco G., Goncalves E., Beaver C.M., Migliardi G., Santos R., Rao Y., Sassi F., Pinnelli M., Ansari R., Harper S., Jackson D.A., McRae R., Pooley R., Wilkinson P., van der Meer D.J., Dow D., Buser-Doepner C.A., Bertotti A., Trusolino L., Stronach E.A., Saez-Rodriguez J., Yusa K., Garnett M.J.
Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens.
Nature 568:511-516(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3; PMCID=PMC6697103
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. 3rd, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=31395879; DOI=10.1038/s41467-019-11415-2; PMCID=PMC6687785
Yu K., Chen B., Aran D., Charalel J., Yau C., Wolf D.M., van 't Veer L.J., Butte A.J., Goldstein T., Sirota M.
Comprehensive transcriptomic analysis of cell lines as models of primary tumors across 22 tumor types.
Nat. Commun. 10:3574.1-3574.11(2019)"