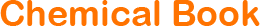
"HT-29人结肠癌细胞代次低|培养基|送STR图谱
传代比例:1:2-1:4(首次传代建议1:2)
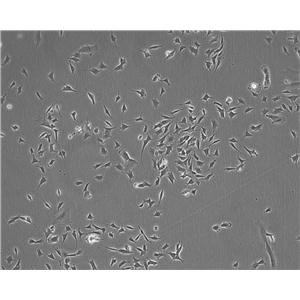
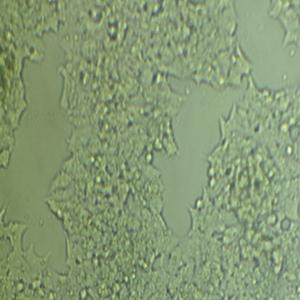
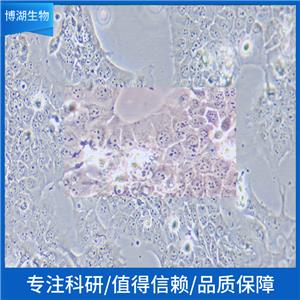
生长特性:贴壁生长
公司细胞系形态漂亮、增殖倍数高、纯度高、功能性强,细胞培养就跟养孩子一个样。养孩子要喂奶,养细胞要加补液,都需要在前期补充足够的营养,初始状态的细胞或刚刚复苏的细胞还要适量加入血清或细胞因子来帮助它们的存活增殖,如果营养物质缺乏,细胞就会不生长甚至死亡。养孩子要从小培养学习,养细胞也得培养宝宝顺利生下来,你会经常抚摸他,给他看各种颜色,刺激他的五感。细胞也是一样,分离后的细胞需要使用特定的细胞因子进行活化、增殖。另外加入因子的种类、因子的浓度、加入时间、加入顺序都会影响细胞最终的结果。养孩子最怕孩子生病,养细胞最怕被污染,平时你会仔细观察宝宝是否呕吐、是否突然哭闹,猜测宝宝是否生病了。对于细胞,我们也需要时刻进行观察的,假如培养液浑浊(污染了),则需要换液后加抗生素;假如细胞增殖不明显,形态变差,则可能是因为营养不足了,对贴壁细胞可以消化后重新用新的培养基接种并加倍加入细胞因子含量;对悬浮细胞增殖能力不强的,则不着急补液,只是先补加血清、细胞因子看是否可以好转。培养时还得全程在无菌的环境,一个小小的偏差,细胞就会死亡。
换液周期:每周2-3次
Potorous tridactylus Kidney 2 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:KYSE30细胞、RCC 7860细胞、Farage细胞
OE33 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:HPMC细胞、Ramos 2G6 4C10细胞、SK Hep1细胞
CWR22Rv1 Cells;背景说明:22RV1是来自异种移植(在阉割引起前列腺癌衰退又在其父亲的雄性激素信赖型CWR22嫁接后复发的小鼠中连续传代)的人前列腺癌上皮细胞系。此细胞系表达前列腺特异抗原。二羟基睾丸脂酮轻微刺激细胞生长,经westernblot检测溶解产物与抗雄性激素受体抗体起免疫反应。EGF刺激细胞生长,但TGFβ-1不能抑制细胞生长。该细胞在裸鼠中成瘤。;传代方法:消化3-5分钟。1:2。3天内可长满。;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:NCI-H2110细胞、Det 562细胞、HME-1细胞
HT-29人结肠癌细胞代次低|培养基|送STR图谱
背景信息:该细胞是1964年由FoghJ用移植培养方法和含15%FBS的F12培养从原发性肿瘤分离的。近来,已建株的培养细胞用含血清的McCoy's5a培养基培养。该细胞系在裸鼠中成瘤,也能在类固醇处理的地鼠中成瘤。该细胞可合成IgA、A、TGFβ结合蛋白和黏素;表达尿激酶受体,但没有检测到血浆酶原活性;不表达CD4,但细胞表面表达半乳糖神经酰胺(HIV的可能替代受体)。该细胞系癌基因c-myc、K-ras、H-ras、N-ras、Myb、sis、fos阳性;p53基因过表达,并且在273位密码子处发
┈订┈购(技术服务)┈热┈线:1┈3┈6┈4┈1┈9┈3┈0┈7┈9┈1【微信同号】┈Q┈Q:3┈1┈8┈0┈8┈0┈7┈3┈2┈4;
DSMZ菌株保藏中心成立于1969年,是德国的国家菌种保藏中心。该中心一直致力于细菌、真菌、质粒、抗菌素、人体和动物细胞、植物病毒等的分类、鉴定和保藏工作。DSMZ菌种保藏中心是欧洲规模最大的生物资源中心,保藏有动物细胞500多株。Riken BRC成立于1920年,是英国的国家菌种保藏中心。该中心一直致力于细菌、真菌、植物病毒等的分类、鉴定和保藏工作。日本Riken BRC(Riken生物资源保藏中心)是全球三大典型培养物收集中心之一。Riken保藏中心提供了很多细胞系。在世界范围内,这些细胞系,都在医学、科学和兽医中具有重要意义。Riken生物资源中心支持了各种学术、健康、食品和兽医机构的研究工作,并在世界各地不同组织的微生物实验室和研究机构中使用。
产品包装:复苏发货:T25培养瓶(一瓶)或冻存发货:1ml冻存管(两支)
来源说明:细胞主要来源ATCC、ECACC、DSMZ、RIKEN等细胞库
HT-29人结肠癌细胞代次低|培养基|送STR图谱
物种来源:人源、鼠源等其它物种来源
NCC-IT Cells;背景说明:详见相关文献介绍;传代方法:1:4—1:8传代,每周换液2—3次;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:NCI-H2198细胞、MGHU1细胞、SUP-T1细胞
V 79 Cells;背景说明:肺;自发永生;雄性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:Liver-02细胞、LS 123细胞、Microbiological Associates-104细胞
NCIH1651 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:8传代,每周换液2次;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:KG-1细胞、HuT 78细胞、NCIH1792细胞
HNE-1 Cells;背景说明:鼻咽部;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:SP-2细胞、DMS 79细胞、NCI-SNU-119细胞
┈订┈购(技术服务)┈热┈线:1┈3┈6┈4┈1┈9┈3┈0┈7┈9┈1【微信同号】┈Q┈Q:3┈1┈8┈0┈8┈0┈7┈3┈2┈4;
形态特性:上皮细胞样
细胞复苏后贴壁细胞较少的问题分析:总结1:复苏过程没有问题,是否是从拿出直接放入温水,还有培养箱,二氧化碳浓度,培养基、PH值等环节。要么加GAO浓度FBS 15-20%,看看能否帮助贴壁,当然也需要考虑血清问题,还有确信拿来的细胞没问题。总结2:首先应该怀疑冻存,实际上复苏出问题的可能非常小,因为操作简单,而且死板。1、你冻存的时候是不是消化的时间过长,这是一般人所注意不到的,即使书上也不讲这个问题,太长的消化时间会让细胞复苏时失去贴壁能力,表现为先贴后死,原因是在你复苏的时候细胞已进入凋亡程序,不可逆转的死亡。2、你的冻存HAO不HAO,是什么,甘油还是DMSO,质量非常重要,否则也会死亡。3、你的冻存的量加的是不是太多,AC推荐是不超过7%,大于5%,太多也不HAO。4、你在冻存的时候是不是把DMSO混均匀,这个有一些影响,但不算太大。5、你的冻存是否按部就班,就是所温度梯度是不是把握严格,很多人容易忘却这个事情,因为这个东西流程长。6、如果你细胞污染,你是否能很快看到,我比我的导师能早一天看到污染。从这个角度讲建议去除离心这步。7、你的细胞在冻存前是否过密。还有,不赞成孵箱污染这个概念的,所有在一个孵箱里的细胞都污染一个细菌的话,这个细菌是源于孵箱的,但这不代表孵箱污染,因为孵箱无论你如何处理都有大量的细菌,问题在操作。每次污染的原因都要尽可能的找,以后就不犯同样的问题,这个很重要,不能靠猜,否则你就有可能细胞养绝Zui后换课题,这个见得太多了,别不当会事。
EFM-192A Cells;背景说明:乳腺癌;胸腔积液转移;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:H524细胞、Anip 973细胞、Mv1Lu细胞
SUM52 Cells;背景说明:乳腺癌;胸腔积液转移;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:PC-12细胞、NCIH2081细胞、NCI-H524细胞
SUDHL4 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:淋巴母细胞;相关产品有:3T3-Swiss albino细胞、MPC-11细胞、GM0637细胞
NTERA-2 Cells;背景说明:畸胎瘤;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:TCMK1细胞、FD-LSC-1细胞、NCI-H378细胞
SK-N-F1 Cells;背景说明:详见相关文献介绍;传代方法:1:4传代,每周换液2次;生长特性:贴壁生长;形态特性:上皮样;相关产品有:Hs 274.T细胞、IM-95细胞、SW 1271细胞
H-220 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Cor L51细胞、SV-HUC-1细胞、PK 15细胞
RM1 Cells;背景说明:前列腺癌;C57BL/6;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:NCIH208细胞、RAMSCs细胞、CFPAC-1细胞
OVCAR-420 Cells;背景说明:卵巢癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:LICR-HN-6细胞、MDA435细胞、SNU-216细胞
LLC-PK1 Cells;背景说明:肾;自发永生;Hampshire;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:TR146细胞、RBE4细胞、A549/ATCC细胞
HCT 8 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:KNS42细胞、P-388D1细胞、WEHI 231细胞
Hu-P-T4 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:COV362细胞、697细胞、Menschliche Und Tierische Zellkulture-3细胞
TK-10 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代;2-3天换液1次。;生长特性:贴壁生长;形态特性:上皮样;相关产品有:Panc 03.27细胞、FLC-7细胞、Hep2细胞
NCIH2172 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代;每周换液2-3次。;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:MCA-205细胞、KY-50细胞、HBMEC细胞
C3H10T1/2CL8 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:SKNFI细胞、TE-15细胞、H-2073细胞
MDA-MB436 Cells;背景说明:该细胞源于一名43岁患有乳腺腺癌女性的胸腔积液。;传代方法:1:2传代,每周换液2—3次;生长特性:贴壁生长;形态特性:多角形;相关产品有:LA-795细胞、X63Ag8.653细胞、Colon 38细胞
VP 229 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:COLO-680N细胞、BTI-Tn-5B1-4细胞、UM-UC3细胞
SK 1 Cells;背景说明:详见相关文献介绍;传代方法:维持细胞浓度在1×105-2×105,每周补液2-3次。;生长特性:悬浮生长;形态特性:球形的;相关产品有:Colo-206F细胞、SW 756细胞、Hep 3B2细胞
Abcam HEK293T EIF2AK4 KO 1 Cells(提供STR鉴定图谱)
AG09980 Cells(提供STR鉴定图谱)
BayGenomics ES cell line NPX373 Cells(提供STR鉴定图谱)
BayGenomics ES cell line XC597 Cells(提供STR鉴定图谱)
BT-L Cells(提供STR鉴定图谱)
COST Cells(提供STR鉴定图谱)
DA04061 Cells(提供STR鉴定图谱)
DA05931 Cells(提供STR鉴定图谱)
GM00298 Cells(提供STR鉴定图谱)
JVM-2 Cells;背景说明:该细胞是 J.V.Melo从一位63岁患有套细胞淋巴瘤白人女性外周血中分离建立的,经EBV介导获得永生化,该细胞表达p16和细胞周期蛋白D2,低水平表达细胞周期蛋白D1。;传代方法:1:3传代,2-3天传一代;生长特性:悬浮生长;形态特性:淋巴母细胞样;相关产品有:3T3-L1 ad细胞、Caco-2 BBe细胞、SaOS-2细胞
HT-29人结肠癌细胞代次低|培养基|送STR图谱
MRC-5 Cells;背景说明:MRC-5细胞系来自14周龄男性胎儿的正常肺组织,该细胞老化前能传代42~46个倍增时间。;传代方法:1:2-1:5传代;每周1-2次。;生长特性:贴壁生长;形态特性:成纤维细胞样;相关产品有:Huh7.5细胞、SW 780细胞、ST2细胞
FF-WT-BJ Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:成纤维细胞样;相关产品有:hTERT-HPNE细胞、R 2 C细胞、H-4细胞
SW-1463 Cells;背景说明:详见相关文献介绍;传代方法:1:3—1:8传代,每周换液1-2次;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:HO1-N-1细胞、MCF.7细胞、BEAS2B细胞
Walker256-TC Cells;背景说明:乳腺癌;雌性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:WM239细胞、SW-954细胞、HN4细胞
IMR 32 Cells;背景说明:该细胞是1967年4月由NicholsWW,LeeJ和DwightS建立,来源于一名13月龄白人男婴腹部肿块,临床诊断为神经母细胞瘤,伴有极少部位的类器官样分化。;传代方法:1:2传代;生长特性:贴壁生长;形态特性:存在两种细胞类型,小的神经母细胞样细胞和大的透明成纤维样细胞;相关产品有:RenCa细胞、MDA-MB-134 VI细胞、H1048细胞
28F11 Cells(提供STR鉴定图谱)
Ca 9-22 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:Caco-2 BBe细胞、CHL细胞、CHP212细胞
H-322 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:LOU-NH91细胞、NCI-H125细胞、KLM1细胞
NCIH2030 Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:4传代;每周换液2-3次。;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:DoHH-2细胞、RH-30细胞、GP2-293细胞
MT-4 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:悬浮生长;形态特性:淋巴母细胞样;圆形;相关产品有:HPAC细胞、PANC0813细胞、BBE细胞
JOSK-M Cells;背景说明:急性单核细胞白血病;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:SKNO-1细胞、Colo699细胞、HcaF细胞
CRFK Cells;背景说明:肾;自发永生;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:TCC-PAN2细胞、JVM3细胞、NCI.H522细胞
HPAF-II Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:Reh细胞、DMS114细胞、A375-SM细胞
Asian Medical Center-Head and Neck cancer-8 Cells;背景说明:喉癌;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:L 540细胞、RMC细胞、P3-X63-Ag8-653细胞
GM28012 Cells(提供STR鉴定图谱)
HAP1 PARP6 (-) 3 Cells(提供STR鉴定图谱)
Mo7e Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:H-2196细胞、CAKI 2细胞、NCI-H510细胞
NCIH2009 Cells;背景说明:详见相关文献介绍;传代方法:每周换液2-3次。;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:FD-LSC-1细胞、CCD1095Sk细胞、CT26.WT细胞
RAMOS2G64C10 Cells;背景说明:详见相关文献介绍;传代方法: 维持细胞浓度在2×105/ml-1×106/ml;根据细胞浓度每2-3天补液1次。;生长特性:悬浮生长 ;形态特性:淋巴母细胞样;相关产品有:McCoy细胞、MADB106细胞、U-87MG细胞
Rat-1 Cells;背景说明:成纤维细胞;自发永生;Fischer 344;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:GM00215A细胞、MDA231细胞、KU-812-F细胞
Ketr3 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:HRA19细胞、VE细胞、RPMI7666细胞
H647ell Cells;背景说明:详见相关文献介绍;传代方法:1:3-1:6传代;每周换液2次。;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:SMMC-7721细胞、C3H10T1/2细胞、OCIAML4细胞
624 Cells;背景说明:黑色素瘤;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:MCMEC细胞、D10.G4.1细胞、CMT 64细胞
HCC-1588 Cells;背景说明:肺鳞癌;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:H-2087细胞、ARH77细胞、KU 19-19细胞
HS-24 Cells(提供STR鉴定图谱)
KOLF2.1J GBA1 D448V SNV/SNV Cells(提供STR鉴定图谱)
MMPR-11 Cells(提供STR鉴定图谱)
NUIGi033-A Cells(提供STR鉴定图谱)
RG-201 Cells(提供STR鉴定图谱)
Ubigene HCT 116 YAP1 KO Cells(提供STR鉴定图谱)
WBD001816 Cells(提供STR鉴定图谱)
HG02056 Cells(提供STR鉴定图谱)
MCF 7B Cells;背景说明:浸润性导管癌;胸腔积液转移;女性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:J-111细胞、CCD 1112SK细胞、Hep 2细胞
Leukemia L1210 Cells;背景说明:该细胞源于用0.2%甲基胆蒽(溶解)涂抹雌性小鼠的皮肤诱发的肿瘤,鼠痘病毒阴性。;传代方法:1:2传代;生长特性:悬浮生长;形态特性:淋巴母细胞样;相关产品有:Normal Rat, August 3, 1983细胞、NCI.H226细胞、HCGC细胞
A2780/Taxol Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:RT112细胞、OVMANA细胞、HSC/mHSC细胞
Pa017C Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:PLA802细胞、CWR22Rv1细胞、Li-7细胞
b.End3 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Calu 6细胞、C1498细胞、8305-C细胞
NWA Cells;背景说明:这株细胞有EB病毒基因组。;传代方法:1:2传代。3天内可长满。;生长特性:悬浮生长 ;形态特性:淋巴母细胞样;相关产品有:NCI H2106细胞、OCI AML3细胞、Natural Killer-92细胞
Swiss 3T3 Cells;背景说明:3T3细胞株是1962年Todaro G和Green H从分离的瑞士小鼠胚胎中建立的;该细胞的生长受接触性抑制,汇合状态的单层细胞密度为40000个细胞/平方厘米;检测结果显示该细胞鼠痘病毒阴性;在中生长较好,在某些玻璃表面上可能状态不佳;细胞生长饱和时其密度可以达到约50000 cells/cm2。;传代方法:1:3传代;3-4天1次。;生长特性:贴壁生长;形态特性:成纤维细胞样;相关产品有:JROECL 19细胞、624细胞、HNE2细胞
SU-DHL6 Cells;背景说明:详见相关文献介绍;传代方法:1:3—1:6传代,3—4天换液1次;生长特性:悬浮生长 ;形态特性:淋巴母细胞样;相关产品有:HuT78细胞、NCI-H661细胞、IOSE-29细胞
SK-N-BE-2 Cells;背景说明:1972年11月从一们多次化疗及放疗的扩散性神经母细胞瘤患儿骨髓穿刺物中建立了SK-N-BE(2)神经母细胞瘤细胞株。 该细胞显示中等水平的多巴胺-β-羟基酶活性。 有报道称SK-N-BE(2)细胞的饱和浓度超过1x106细胞/平方厘米。细胞形态多样,有的有长突触,有的呈上皮细胞样。 细胞会聚集,形成团块并浮起;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮细胞样;相关产品有:ID8/MOSEC细胞、BpRc1细胞、SK-CO-1细胞
CCRF/CEM Cells;背景说明:G.E. Foley 等人建立了类淋巴母细胞细胞株CCRF-CEM。 细胞是1964年11月从一位四岁白人女性急性淋巴细胞白血病患者的外周血白血球衣中得到。此细胞系从香港收集而来。;传代方法:1:2传代。3天内可长满。;生长特性:悬浮生长;形态特性:淋巴母细胞样;相关产品有:M-1细胞、PL 5细胞、DHBE细胞
OAW-42 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:SKNDZ细胞、Granta519细胞、NCIH2085细胞
OCI/AML-2 Cells;背景说明:急性髓系白血病;男性;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:悬浮;形态特性:详见产品说明书;相关产品有:HCC94细胞、SKNFI细胞、HC11细胞
SF126 Cells;背景说明:该细胞来源于星形胶质细胞瘤;胶质纤维酸性蛋白(GFAP)阴性;可以特异地结合β-内啡肽。;传代方法:1:3传代;3-4天1次。;生长特性:贴壁生长;形态特性:成纤维细胞;相关产品有:KMH-2细胞、MCA 205细胞、MKN 7细胞
B16-F10 Cells;背景说明:B16-F10是B16-F0的亚系。;传代方法:消化3-5分钟。1:2。3天内可长满。;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:Sp2/mIL-6细胞、HEK293-A细胞、EU-4细胞
U-343-MG Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长 ;形态特性:详见产品说明书;相关产品有:OUMS-27细胞、CMT 93细胞、H1568细胞
STBCi283-A Cells(提供STR鉴定图谱)
ECC12 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:HEK-EBNA细胞、U-118 MG细胞、Tadarida brasiliensis 1 lung细胞
S91 Cells;背景说明:黑色素瘤;雄性;DBA;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁;形态特性:详见产品说明书;相关产品有:EFM192细胞、Walker-256细胞、MDAMB134细胞
Hs 863.T Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代;每周换液2-3次。;生长特性:贴壁生长;形态特性:成纤维细胞;相关产品有:CHO-Lec1细胞、A431细胞、WEHI-231细胞
PL11 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:BHK细胞、Madin Darby Canine Kidney细胞、PF-382细胞
MIN6 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:详见产品说明书;相关产品有:NCIH1568细胞、10RGB细胞、LS 123细胞
AX-Mel Cells;背景说明:详见相关文献介绍;传代方法:1:4-1:6传代,2-3天换液1次。;生长特性:贴壁生长;形态特性:上皮细胞;相关产品有:TCMK-1细胞、OCUM-1细胞、AE 1201细胞
HT-29人结肠癌细胞代次低|培养基|送STR图谱
SNU-423 Cells;背景说明:详见相关文献介绍;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:CNE-2细胞、SRA01/04 (HLE)细胞、CTLA4 Ig-24细胞
U-118-MG Cells;背景说明:注意: 据报道来自不同个体的胶质母细胞瘤细胞株U-118 MG (HTB-15) 和 U-138 MG (HTB-16)有着一致的VNTR和相近的STR模式。 U-118 MG 和 U-138 MG细胞遗传学上很相似并有至少六个衍生标记染色体。 这是1966年至1969年间J. Ponten和同事从恶性神经胶质瘤中构建的细胞株中的一株(其它包括ATCC HTB-14和 ATCC HTB-16 and ATCC HTB-17)。 1987年用BM-Cycline培养6周去除了支原体污染。 ;传代方法: 消化3-5分钟。1:2传代。3天内可长满。;生长特性:贴壁生长;形态特性:混合型;相关产品有:KP4细胞、Hs611T细胞、NB-9细胞
STTG1 Cells;背景说明:详见相关文献介绍;传代方法:1:3—1:6传代;每周换液2-3次;生长特性:贴壁生长;形态特性:星形胶质细胞;相关产品有:QGY 7701细胞、LNCaP-Clone-FGC细胞、SKRC20细胞
Capan-2 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:4传代,2-3天换液1次。;生长特性:贴壁生长;形态特性:多边形;相关产品有:H-1341细胞、Hopkins-92细胞、WEHI-3B细胞
HBSMC Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:Hs 832(C).T细胞、HC11 Mammary Epithelium细胞、EL4细胞
SLMT-1 Cells;背景说明:详见相关文献介绍;传代方法:1:2-1:3传代;每周换液2-3次。;生长特性:贴壁或悬浮,详见产品说明书部分;形态特性:详见产品说明书;相关产品有:L5178Y TK+/- (clone 3.7.2C)细胞、SW954细胞、Walker-256-TC细胞
HBL-100 Cells;背景说明:该细胞由E.V.Gaffney及其同事从一位没有乳癌家族史的供者乳汁中建立,培养出来的细胞染色体组型在第7代时就不正常;电镜照片显示有微丝、张力原纤维和桥粒;Southern转移表明有整合型SV40病毒基因,当作正常细胞。;传代方法:1:2传代;生长特性:贴壁生长;形态特性:上皮样;相关产品有:Instituto Biologico-Rim Suino-2细胞、BT-20细胞、H-2170细胞
Hs819.T Cells;背景说明:详见相关文献介绍;传代方法:1:2—1:3传代;每周换液2-3次;生长特性:贴壁生长;形态特性:成纤维;相关产品有:CBRH7919细胞、SF 126细胞、786.O细胞
BayGenomics ES cell line RRO624 Cells(提供STR鉴定图谱)
BayGenomics ES cell line YHC102 Cells(提供STR鉴定图谱)
HCD-57 SHP2-E76K Cells(提供STR鉴定图谱)
PCRP-CCDC101-1D10 Cells(提供STR鉴定图谱)
BP8 Cells(提供STR鉴定图谱)
HPS0320 Cells(提供STR鉴定图谱)
" "PubMed=327080; DOI=10.1093/jnci/59.1.221
Fogh J., Fogh J.M., Orfeo T.
One hundred and twenty-seven cultured human tumor cell lines producing tumors in nude mice.
J. Natl. Cancer Inst. 59:221-226(1977)
PubMed=833871; DOI=10.1093/jnci/58.2.209
Fogh J., Wright W.C., Loveless J.D.
Absence of HeLa cell contamination in 169 cell lines derived from human tumors.
J. Natl. Cancer Inst. 58:209-214(1977)
PubMed=864752; DOI=10.1093/jnci/58.6.1743
Marshall C.J., Franks L.M., Carbonell A.W.
Markers of neoplastic transformation in epithelial cell lines derived from human carcinomas.
J. Natl. Cancer Inst. 58:1743-1751(1977)
PubMed=77569; DOI=10.1111/j.1399-0039.1978.tb01259.x
Espmark J.A., Ahlqvist-Roth L., Sarne L., Persson A.
Tissue typing of cells in culture. III. HLA antigens of established human cell lines. Attempts at typing by the mixed hemadsorption technique.
Tissue Antigens 11:279-286(1978)
PubMed=626984
Erickson L.C., Osieka R., Kohn K.W.
Differential repair of 1-(2-chloroethyl)-3-(4-methylcyclohexyl)-1- nitrosourea-induced DNA damage in two human colon tumor cell lines.
Cancer Res. 38:802-808(1978)
PubMed=468301; PMCID=PMC1457289
Trejdosiewicz L.K., Trejdosiewicz A.J., Ling N.R., Dykes P.W.
Growth enhancing property of human monocytes from normal donors and cancer patients.
Immunology 37:247-252(1979)
PubMed=6256643; DOI=10.1038/288724a0
Day R.S. 3rd, Ziolkowski C.H.J., Scudiero D.A., Meyer S.A., Lubiniecki A.S., Girardi A.J., Galloway S.M., Bynum G.D.
Defective repair of alkylated DNA by human tumour and SV40-transformed human cell strains.
Nature 288:724-727(1980)
PubMed=6445228
Kimball P.M., Brattain M.G.
Isolation of a cellular subpopulation from a human colonic carcinoma cell line.
Cancer Res. 40:1574-1579(1980)
PubMed=22282976; DOI=10.1093/carcin/1.1.21
Day R.S. 3rd, Ziolkowski C.H.J., Scudiero D.A., Meyer S.A., Mattern M.R.
Human tumor cell strains defective in the repair of alkylation damage.
Carcinogenesis 1:21-32(1980)
PubMed=7017212; DOI=10.1093/jnci/66.6.1003
Pollack M.S., Heagney S.D., Livingston P.O., Fogh J.
HLA-A, B, C and DR alloantigen expression on forty-six cultured human tumor cell lines.
J. Natl. Cancer Inst. 66:1003-1012(1981)
PubMed=7459858
Rousset M., Zweibaum A., Fogh J.
Presence of glycogen and growth-related variations in 58 cultured human tumor cell lines of various tissue origins.
Cancer Res. 41:1165-1170(1981)
PubMed=6220172
Dracopoli N.C., Fogh J.
Polymorphic enzyme analysis of cultured human tumor cell lines.
J. Natl. Cancer Inst. 70:469-476(1983)
PubMed=6825208; DOI=10.1093/carcin/4.2.199
Yarosh D.B., Foote R.S., Mitra S., Day R.S. 3rd
Repair of O6-methylguanine in DNA by demethylation is lacking in Mer- human tumor cell strains.
Carcinogenesis 4:199-205(1983)
PubMed=6500159; DOI=10.1159/000163283
Gershwin M.E., Lentz D., Owens R.B.
Relationship between karyotype of tissue culture lines and tumorigenicity in nude mice.
Exp. Cell Biol. 52:361-370(1984)
PubMed=6582512; DOI=10.1073/pnas.81.2.568; PMCID=PMC344720
Mattes M.J., Cordon-Cardo C., Lewis J.L. Jr., Old L.J., Lloyd K.O.
Cell surface antigens of human ovarian and endometrial carcinoma defined by mouse monoclonal antibodies.
Proc. Natl. Acad. Sci. U.S.A. 81:568-572(1984)
PubMed=3518877; DOI=10.3109/07357908609038260
Fogh J.
Human tumor lines for cancer research.
Cancer Invest. 4:157-184(1986)
PubMed=3472642; DOI=10.1016/0165-4608(87)90267-6
Chen T.-R., Drabkowski D.J., Hay R.J., Macy M.L., Peterson W.D. Jr.
WiDr is a derivative of another colon adenocarcinoma cell line, HT-29.
Cancer Genet. Cytogenet. 27:125-134(1987)
PubMed=3335022
Alley M.C., Scudiero D.A., Monks A., Hursey M.L., Czerwinski M.J., Fine D.L., Abbott B.J., Mayo J.G., Shoemaker R.H., Boyd M.R.
Feasibility of drug screening with panels of human tumor cell lines using a microculture tetrazolium assay.
Cancer Res. 48:589-601(1988)
PubMed=3349466
Chantret I., Barbat A., Dussaulx E., Brattain M.G., Zweibaum A.
Epithelial polarity, villin expression, and enterocytic differentiation of cultured human colon carcinoma cells: a survey of twenty cell lines.
Cancer Res. 48:1936-1942(1988)
PubMed=2288877
Hafez M.M., Infante D., Winawer S.J., Friedman E.
Transforming growth factor beta 1 acts as an autocrine-negative growth regulator in colon enterocytic differentiation but not in goblet cell maturation.
Cell Growth Differ. 1:617-626(1990)
PubMed=1778766; DOI=10.1111/j.1349-7006.1991.tb01816.x; PMCID=PMC5918361
Takeshima E., Hamaguchi M., Watanabe T., Akiyama S., Kataoka M., Ohnishi Y., Xiao H.-Y., Nagai Y., Takagi H.
Aberrant elevation of tyrosine-specific phosphorylation in human gastric cancer cells.
Jpn. J. Cancer Res. 82:1428-1435(1991)
PubMed=1937958; DOI=10.1002/ijc.2910490516
Lesuffleur T., Kornowski A., Luccioni C., Muleris M., Barbat A., Beaumatin J., Dussaulx E., Dutrillaux B., Zweibaum A.
Adaptation to 5-fluorouracil of the heterogeneous human colon tumor cell line HT-29 results in the selection of cells committed to differentiation.
Int. J. Cancer 49:721-730(1991)
PubMed=2041050; DOI=10.1093/jnci/83.11.757
Monks A., Scudiero D.A., Skehan P., Shoemaker R.H., Paull K.D., Vistica D.T., Hose C.D., Langley J., Cronise P., Vaigro-Wolff A., Gray-Goodrich M., Campbell H., Mayo J.G., Boyd M.R.
Feasibility of a high-flux anticancer drug screen using a diverse panel of cultured human tumor cell lines.
J. Natl. Cancer Inst. 83:757-766(1991)
PubMed=1389533; DOI=10.1016/0959-8049(92)90031-V
Lahm H., Petral-Malec D., Yilmaz-Ceyhan A., Fischer J.R., Lorenzoni M., Givel J.-C., Odartchenko N.
Growth stimulation of a human colorectal carcinoma cell line by interleukin-1 and -6 and antagonistic effects of transforming growth factor beta 1.
Eur. J. Cancer 28A:1894-1899(1992)
PubMed=1730571; DOI=10.1007/BF02631079
Goodwin T.J., Jessup J.M., Wolf D.A.
Morphologic differentiation of colon carcinoma cell lines HT-29 and HT-29KM in rotating-wall vessels.
In Vitro Cell. Dev. Biol. Anim. 28:47-60(1992)
PubMed=8464898; DOI=10.1073/pnas.90.7.2842; PMCID=PMC46192
Browning M.J., Krausa P., Rowan A.J., Bicknell D.C., Bodmer J.G., Bodmer W.F.
Tissue typing the HLA-A locus from genomic DNA by sequence-specific PCR: comparison of HLA genotype and surface expression on colorectal tumor cell lines.
Proc. Natl. Acad. Sci. U.S.A. 90:2842-2845(1993)
PubMed=7972006; DOI=10.1073/pnas.91.23.11045; PMCID=PMC45163
Okamoto A., Demetrick D.J., Spillare E.A., Hagiwara K., Hussain S.P., Bennett W.P., Forrester K., Gerwin B.I., Serrano M., Beach D.H., Harris C.C.
Mutations and altered expression of p16INK4 in human cancer.
Proc. Natl. Acad. Sci. U.S.A. 91:11045-11049(1994)
PubMed=7651727
Kastrinakis W.V., Ramchurren N., Rieger K.M., Hess D.T., Loda M., Steele G., Summerhayes I.C.
Increased incidence of p53 mutations is associated with hepatic metastasis in colorectal neoplastic progression.
Oncogene 11:647-652(1995)
PubMed=8895552; DOI=10.1002/(SICI)1097-0215(19960927)68:1<126::AID-IJC22>3.0.CO;2-8
Suardet L., Li C., Little J.B.
Radio-induced modulation of transforming growth factor beta1 sensitivity in a p53 wild-type human colorectal-cancer cell line.
Int. J. Cancer 68:126-131(1996)
PubMed=9000147
Cottu P.-H., Muzeau F., Estreicher A., Flejou J.-F., Iggo R.D., Thomas G., Hamelin R.
Inverse correlation between RER+ status and p53 mutation in colorectal cancer cell lines.
Oncogene 13:2727-2730(1996)
PubMed=9000572
Hoang J.-M., Cottu P.-H., Thuille B., Salmon R.J., Thomas G., Hamelin R.
BAT-26, an indicator of the replication error phenotype in colorectal cancers and cell lines.
Cancer Res. 57:300-303(1997)
PubMed=9294210; DOI=10.1073/pnas.94.19.10330; PMCID=PMC23362
Ilyas M., Tomlinson I.P.M., Rowan A.J., Pignatelli M., Bodmer W.F.
Beta-catenin mutations in cell lines established from human colorectal cancers.
Proc. Natl. Acad. Sci. U.S.A. 94:10330-10334(1997)
PubMed=10612807; DOI=10.1002/(SICI)1098-2264(200002)27:2<183::AID-GCC10>3.0.CO;2-P; PMCID=PMC4721570
Ghadimi B.M., Sackett D.L., Difilippantonio M.J., Schrock E., Neumann T., Jauho A., Auer G., Ried T.
Centrosome amplification and instability occurs exclusively in aneuploid, but not in diploid colorectal cancer cell lines, and correlates with numerical chromosomal aberrations.
Genes Chromosomes Cancer 27:183-190(2000)
PubMed=10700174; DOI=10.1038/73432
Ross D.T., Scherf U., Eisen M.B., Perou C.M., Rees C., Spellman P.T., Iyer V.R., Jeffrey S.S., van de Rijn M., Waltham M.C., Pergamenschikov A., Lee J.C.F., Lashkari D., Shalon D., Myers T.G., Weinstein J.N., Botstein D., Brown P.O.
Systematic variation in gene expression patterns in human cancer cell lines.
Nat. Genet. 24:227-235(2000)
PubMed=10737795; DOI=10.1073/pnas.97.7.3352; PMCID=PMC16243
Rowan A.J., Lamlum H., Ilyas M., Wheeler J.M.D., Straub J., Papadopoulou A., Bicknell D.C., Bodmer W.F., Tomlinson I.P.M.
APC mutations in sporadic colorectal tumors: a mutational 'hotspot' and interdependence of the 'two hits'.
Proc. Natl. Acad. Sci. U.S.A. 97:3352-3357(2000)
PubMed=11226274; DOI=10.1073/pnas.041603298; PMCID=PMC30173
Abdel-Rahman W.M., Katsura K., Rens W., Gorman P.A., Sheer D., Bicknell D.C., Bodmer W.F., Arends M.J., Wyllie A.H., Edwards P.A.W.
Spectral karyotyping suggests additional subsets of colorectal cancers characterized by pattern of chromosome rearrangement.
Proc. Natl. Acad. Sci. U.S.A. 98:2538-2543(2001)
PubMed=11414198; DOI=10.1007/s004320000207
Lahm H., Andre S., Hoeflich A., Fischer J.R., Sordat B., Kaltner H., Wolf E., Gabius H.-J.
Comprehensive galectin fingerprinting in a panel of 61 human tumor cell lines by RT-PCR and its implications for diagnostic and therapeutic procedures.
J. Cancer Res. Clin. Oncol. 127:375-386(2001)
PubMed=11416159; DOI=10.1073/pnas.121616198; PMCID=PMC35459
Masters J.R.W., Thomson J.A., Daly-Burns B., Reid Y.A., Dirks W.G., Packer P., Toji L.H., Ohno T., Tanabe H., Arlett C.F., Kelland L.R., Harrison M., Virmani A.K., Ward T.H., Ayres K.L., Debenham P.G.
Short tandem repeat profiling provides an international reference standard for human cell lines.
Proc. Natl. Acad. Sci. U.S.A. 98:8012-8017(2001)
PubMed=11526487; DOI=10.1038/sj.onc.1204611
Gayet J., Zhou X.-P., Duval A., Rolland S., Hoang J.-M., Cottu P.-H., Hamelin R.
Extensive characterization of genetic alterations in a series of human colorectal cancer cell lines.
Oncogene 20:5025-5032(2001)
PubMed=11668190; DOI=10.1177/002215540104901105
Quentmeier H., Osborn M., Reinhardt J., Zaborski M., Drexler H.G.
Immunocytochemical analysis of cell lines derived from solid tumors.
J. Histochem. Cytochem. 49:1369-1378(2001)
PubMed=11687795; DOI=10.1038/ng754
Snijders A.M., Nowak N.J., Segraves R., Blackwood S., Brown N., Conroy J., Hamilton G., Hindle A.K., Huey B., Kimura K., Law S., Myambo K., Palmer J., Ylstra B., Yue J.P., Gray J.W., Jain A.N., Pinkel D., Albertson D.G.
Assembly of microarrays for genome-wide measurement of DNA copy number.
Nat. Genet. 29:263-264(2001)
PubMed=11921276; DOI=10.1002/gcc.10003
Kawai K., Viars C., Arden K.C., Tarin D., Urquidi V., Goodison S.
Comprehensive karyotyping of the HT-29 colon adenocarcinoma cell line.
Genes Chromosomes Cancer 34:1-8(2002)
PubMed=12068308; DOI=10.1038/nature00766
Davies H.R., Bignell G.R., Cox C., Stephens P.J., Edkins S., Clegg S., Teague J.W., Woffendin H., Garnett M.J., Bottomley W., Davis N., Dicks E., Ewing R., Floyd Y., Gray K., Hall S., Hawes R., Hughes J., Kosmidou V., Menzies A., Mould C., Parker A., Stevens C., Watt S., Hooper S., Wilson R., Jayatilake H., Gusterson B.A., Cooper C.S., Shipley J.M., Hargrave D., Pritchard-Jones K., Maitland N.J., Chenevix-Trench G., Riggins G.J., Bigner D.D., Palmieri G., Cossu A., Flanagan A.M., Nicholson A., Ho J.W.C., Leung S.Y., Yuen S.T., Weber B.L., Seigler H.F., Darrow T.L., Paterson H.F., Marais R., Marshall C.J., Wooster R., Stratton M.R., Futreal P.A.
Mutations of the BRAF gene in human cancer.
Nature 417:949-954(2002)
PubMed=15748285; DOI=10.1186/1479-5876-3-11; PMCID=PMC555742
Adams S., Robbins F.-M., Chen D., Wagage D., Holbeck S.L., Morse H.C. 3rd, Stroncek D., Marincola F.M.
HLA class I and II genotype of the NCI-60 cell lines.
J. Transl. Med. 3:11.1-11.8(2005)
PubMed=15900046; DOI=10.1093/jnci/dji133
Mashima T., Oh-hara T., Sato S., Mochizuki M., Sugimoto Y., Yamazaki K., Hamada J.-i., Tada M., Moriuchi T., Ishikawa Y., Kato Y., Tomoda H., Yamori T., Tsuruo T.
p53-defective tumors with a functional apoptosome-mediated pathway: a new therapeutic target.
J. Natl. Cancer Inst. 97:765-777(2005)
PubMed=16083285; DOI=10.1021/pr050048h
Kim J.-E., Tannenbaum S.R., White F.M.
Global phosphoproteome of HT-29 human colon adenocarcinoma cells.
J. Proteome Res. 4:1339-1346(2005)
PubMed=17088437; DOI=10.1158/1535-7163.MCT-06-0433; PMCID=PMC2705832
Ikediobi O.N., Davies H.R., Bignell G.R., Edkins S., Stevens C., O'Meara S., Santarius T., Avis T., Barthorpe S., Brackenbury L., Buck G., Butler A.P., Clements J., Cole J., Dicks E., Forbes S., Gray K., Halliday K., Harrison R., Hills K., Hinton J., Hunter C., Jenkinson A., Jones D., Kosmidou V., Lugg R., Menzies A., Miroo T., Parker A., Perry J., Raine K.M., Richardson D., Shepherd R., Small A., Smith R., Solomon H., Stephens P.J., Teague J.W., Tofts C., Varian J., Webb T., West S., Widaa S., Yates A., Reinhold W.C., Weinstein J.N., Stratton M.R., Futreal P.A., Wooster R.
Mutation analysis of 24 known cancer genes in the NCI-60 cell line set.
Mol. Cancer Ther. 5:2606-2612(2006)
PubMed=18258742; DOI=10.1073/pnas.0712176105; PMCID=PMC2268141
Emaduddin M., Bicknell D.C., Bodmer W.F., Feller S.M.
Cell growth, global phosphotyrosine elevation, and c-Met phosphorylation through Src family kinases in colorectal cancer cells.
Proc. Natl. Acad. Sci. U.S.A. 105:2358-2362(2008)
PubMed=19372543; DOI=10.1158/1535-7163.MCT-08-0921; PMCID=PMC4020356
Lorenzi P.L., Reinhold W.C., Varma S., Hutchinson A.A., Pommier Y., Chanock S.J., Weinstein J.N.
DNA fingerprinting of the NCI-60 cell line panel.
Mol. Cancer Ther. 8:713-724(2009)
PubMed=19927377; DOI=10.1002/gcc.20730; PMCID=PMC2818350
Knutsen T., Padilla-Nash H.M., Wangsa D., Barenboim-Stapleton L., Camps J., McNeil N.E., Difilippantonio M.J., Ried T.
Definitive molecular cytogenetic characterization of 15 colorectal cancer cell lines.
Genes Chromosomes Cancer 49:204-223(2010)
PubMed=19941903; DOI=10.1016/j.jviromet.2009.11.022
Karger A., Bettin B., Lenk M., Mettenleiter T.C.
Rapid characterisation of cell cultures by matrix-assisted laser desorption/ionisation mass spectrometric typing.
J. Virol. Methods 164:116-121(2010)
PubMed=20164919; DOI=10.1038/nature08768; PMCID=PMC3145113
Bignell G.R., Greenman C.D., Davies H.R., Butler A.P., Edkins S., Andrews J.M., Buck G., Chen L., Beare D., Latimer C., Widaa S., Hinton J., Fahey C., Fu B.-Y., Swamy S., Dalgliesh G.L., Teh B.T., Deloukas P., Yang F.-T., Campbell P.J., Futreal P.A., Stratton M.R.
Signatures of mutation and selection in the cancer genome.
Nature 463:893-898(2010)
PubMed=20215515; DOI=10.1158/0008-5472.CAN-09-3458; PMCID=PMC2881662
Rothenberg S.M., Mohapatra G., Rivera M.N., Winokur D., Greninger P., Nitta M., Sadow P.M., Sooriyakumar G., Brannigan B.W., Ulman M.J., Perera R.M., Wang R., Tam A., Ma X.-J., Erlander M., Sgroi D.C., Rocco J.W., Lingen M.W., Cohen E.E.W., Louis D.N., Settleman J., Haber D.A.
A genome-wide screen for microdeletions reveals disruption of polarity complex genes in diverse human cancers.
Cancer Res. 70:2158-2164(2010)
PubMed=20570890; DOI=10.1158/0008-5472.CAN-10-0192; PMCID=PMC2943514
Janakiraman M., Vakiani E., Zeng Z.-S., Pratilas C.A., Taylor B.S., Chitale D., Halilovic E., Wilson M., Huberman K., Ricarte Filho J.C.M., Persaud Y., Levine D.A., Fagin J.A., Jhanwar S.C., Mariadason J.M., Lash A., Ladanyi M., Saltz L.B., Heguy A., Paty P.B., Solit D.B.
Genomic and biological characterization of exon 4 KRAS mutations in human cancer.
Cancer Res. 70:5901-5911(2010)
PubMed=20606684; DOI=10.1038/sj.bjc.6605780; PMCID=PMC2920028
Bracht K., Nicholls A.M., Liu Y., Bodmer W.F.
5-fluorouracil response in a large panel of colorectal cancer cell lines is associated with mismatch repair deficiency.
Br. J. Cancer 103:340-346(2010)
PubMed=20831567; DOI=10.1111/j.1582-4934.2010.01170.x; PMCID=PMC3918049
Ma Y.-L., Zhang P., Wang F., Moyer M.P., Yang J.-J., Liu Z.-H., Peng J.-Y., Chen H.-Q., Zhou Y.-K., Liu W.-J., Qin H.-L.
Human embryonic stem cells and metastatic colorectal cancer cells shared the common endogenous human microRNA-26b.
J. Cell. Mol. Med. 15:1941-1954(2011)
PubMed=22068913; DOI=10.1073/pnas.1111840108; PMCID=PMC3219108
Gillet J.-P., Calcagno A.M., Varma S., Marino M., Green L.J., Vora M.I., Patel C., Orina J.N., Eliseeva T.A., Singal V., Padmanabhan R., Davidson B., Ganapathi R., Sood A.K., Rueda B.R., Ambudkar S.V., Gottesman M.M.
Redefining the relevance of established cancer cell lines to the study of mechanisms of clinical anti-cancer drug resistance.
Proc. Natl. Acad. Sci. U.S.A. 108:18708-18713(2011)
PubMed=21912889; DOI=10.1007/s10637-011-9744-z
Sutherland H.S., Hwang I.Y., Marshall E.S., Lindsay B.S., Denny W.A., Gilchrist C., Joseph W.R., Greenhalgh D., Richardson E., Kestell P., Ding A., Baguley B.C.
Therapeutic reactivation of mutant p53 protein by quinazoline derivatives.
Invest. New Drugs 30:2035-2045(2012)
PubMed=22336246; DOI=10.1016/j.bmc.2012.01.017
Kong D.-X., Yamori T.
JFCR39, a panel of 39 human cancer cell lines, and its application in the discovery and development of anticancer drugs.
Bioorg. Med. Chem. 20:1947-1951(2012)
PubMed=22347499; DOI=10.1371/journal.pone.0031628; PMCID=PMC3276511
Ruan X.-Y., Kocher J.-P.A., Pommier Y., Liu H.-F., Reinhold W.C.
Mass homozygotes accumulation in the NCI-60 cancer cell lines as compared to HapMap trios, and relation to fragile site location.
PLoS ONE 7:E31628-E31628(2012)
PubMed=22384151; DOI=10.1371/journal.pone.0032096; PMCID=PMC3285665
Lee J.-S., Kim Y.K., Kim H.J., Hajar S., Tan Y.L., Kang N.-Y., Ng S.H., Yoon C.N., Chang Y.-T.
Identification of cancer cell-line origins using fluorescence image-based phenomic screening.
PLoS ONE 7:E32096-E32096(2012)
PubMed=22460905; DOI=10.1038/nature11003; PMCID=PMC3320027
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=22628656; DOI=10.1126/science.1218595; PMCID=PMC3526189
Jain M., Nilsson R., Sharma S., Madhusudhan N., Kitami T., Souza A.L., Kafri R., Kirschner M.W., Clish C.B., Mootha V.K.
Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation.
Science 336:1040-1044(2012)
PubMed=23272949; DOI=10.1186/1755-8794-5-66; PMCID=PMC3543849
Schlicker A., Beran G., Chresta C.M., McWalter G., Pritchard A., Weston S., Runswick S., Davenport S., Heathcote K., Castro D.A., Orphanides G., French T., Wessels L.F.A.
Subtypes of primary colorectal tumors correlate with response to targeted treatment in colorectal cell lines.
BMC Med. Genomics 5:66.1-66.15(2012)
PubMed=22940288; DOI=10.1016/j.jsbmb.2012.08.003; PMCID=PMC3695570
Hobaus J., Fetahu I.S., Khorchide M., Manhardt T., Kallay E.
Epigenetic regulation of the 1,25-dihydroxyvitamin D3 24-hydroxylase (CYP24A1) in colon cancer cells.
J. Steroid Biochem. Mol. Biol. 136:296-299(2013)
PubMed=23546019; DOI=10.3892/ijo.2013.1868
Petitprez A., Poindessous V., Ouaret D., Regairaz M., Bastian G., Guerin E., Escargueil A.E., Larsen A.K.
Acquired irinotecan resistance is accompanied by stable modifications of cell cycle dynamics independent of MSI status.
Int. J. Oncol. 42:1644-1653(2013)
PubMed=23631600; DOI=10.1021/pr400260h
Loftus N.J., Lai L., Wilkinson R.W., Odedra R., Wilson I.D., Barnes A.J.
Global metabolite profiling of human colorectal cancer xenografts in mice using HPLC-MS/MS.
J. Proteome Res. 12:2980-2986(2013)
PubMed=23856246; DOI=10.1158/0008-5472.CAN-12-3342; PMCID=PMC4893961
Abaan O.D., Polley E.C., Davis S.R., Zhu Y.-L.J., Bilke S., Walker R.L., Pineda M.A., Gindin Y., Jiang Y., Reinhold W.C., Holbeck S.L., Simon R.M., Doroshow J.H., Pommier Y., Meltzer P.S.
The exomes of the NCI-60 panel: a genomic resource for cancer biology and systems pharmacology.
Cancer Res. 73:4372-4382(2013)
PubMed=23932154; DOI=10.1016/j.radonc.2013.06.032
Salendo J., Spitzner M., Kramer F., Zhang X., Jo P., Wolff H.A., Kitz J., Kaulfuss S., Beissbarth T., belstein M., Ghadimi M., Grade M., Gaedcke J.
Identification of a microRNA expression signature for chemoradiosensitivity of colorectal cancer cells, involving miRNAs-320a, -224, -132 and let7g.
Radiother. Oncol. 108:451-457(2013)
PubMed=23933261; DOI=10.1016/j.celrep.2013.07.018
Moghaddas Gholami A., Hahne H., Wu Z.-X., Auer F.J., Meng C., Wilhelm M., Kuster B.
Global proteome analysis of the NCI-60 cell line panel.
Cell Rep. 4:609-620(2013)
PubMed=24042735; DOI=10.1038/oncsis.2013.35; PMCID=PMC3816225
Ahmed D., Eide P.W., Eilertsen I.A., Danielsen S.A., Eknaes M., Hektoen M., Lind G.E., Lothe R.A.
Epigenetic and genetic features of 24 colon cancer cell lines.
Oncogenesis 2:e71.1-e71.8(2013)
PubMed=24279929; DOI=10.1186/2049-3002-1-20; PMCID=PMC4178206
Dolfi S.C., Chan L.L.-Y., Qiu J., Tedeschi P.M., Bertino J.R., Hirshfield K.M., Oltvai Z.N., Vazquez A.
The metabolic demands of cancer cells are coupled to their size and protein synthesis rates.
Cancer Metab. 1:20.1-20.13(2013)
PubMed=24670534; DOI=10.1371/journal.pone.0092047; PMCID=PMC3966786
Varma S., Pommier Y., Sunshine M., Weinstein J.N., Reinhold W.C.
High resolution copy number variation data in the NCI-60 cancer cell lines from whole genome microarrays accessible through CellMiner.
PLoS ONE 9:E92047-E92047(2014)
PubMed=24755471; DOI=10.1158/0008-5472.CAN-14-0013
Mouradov D., Sloggett C., Jorissen R.N., Love C.G., Li S., Burgess A.W., Arango D., Strausberg R.L., Buchanan D., Wormald S., O'Connor L., Wilding J.L., Bicknell D.C., Tomlinson I.P.M., Bodmer W.F., Mariadason J.M., Sieber O.M.
Colorectal cancer cell lines are representative models of the main molecular subtypes of primary cancer.
Cancer Res. 74:3238-3247(2014)
PubMed=25984343; DOI=10.1038/sdata.2014.35; PMCID=PMC4432652
Cowley G.S., Weir B.A., Vazquez F., Tamayo P., Scott J.A., Rusin S., East-Seletsky A., Ali L.D., Gerath W.F.J., Pantel S.E., Lizotte P.H., Jiang G.-Z., Hsiao J., Tsherniak A., Dwinell E., Aoyama S., Okamoto M., Harrington W., Gelfand E.T., Green T.M., Tomko M.J., Gopal S., Wong T.C., Li H.-B., Howell S., Stransky N., Liefeld T., Jang D., Bistline J., Meyers B.H., Armstrong S.A., Anderson K.C., Stegmaier K., Reich M., Pellman D., Boehm J.S., Mesirov J.P., Golub T.R., Root D.E., Hahn W.C.
Parallel genome-scale loss of function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies.
Sci. Data 1:140035-140035(2014)
PubMed=25485619; DOI=10.1038/nbt.3080
Klijn C., Durinck S., Stawiski E.W., Haverty P.M., Jiang Z.-S., Liu H.-B., Degenhardt J., Mayba O., Gnad F., Liu J.-F., Pau G., Reeder J., Cao Y., Mukhyala K., Selvaraj S.K., Yu M.-M., Zynda G.J., Brauer M.J., Wu T.D., Gentleman R.C., Manning G., Yauch R.L., Bourgon R., Stokoe D., Modrusan Z., Neve R.M., de Sauvage F.J., Settleman J., Seshagiri S., Zhang Z.-M.
A comprehensive transcriptional portrait of human cancer cell lines.
Nat. Biotechnol. 33:306-312(2015)
PubMed=25841592; DOI=10.1016/j.jprot.2015.03.019
Piersma S.R., Knol J.C., de Reus I., Labots M., Sampadi B.K., Pham T.V., Ishihama Y., Verheul H.M.W., Jimenez C.R.
Feasibility of label-free phosphoproteomics and application to base-line signaling of colorectal cancer cell lines.
J. Proteomics 127:247-258(2015)
PubMed=25877200; DOI=10.1038/nature14397
Yu M., Selvaraj S.K., Liang-Chu M.M.Y., Aghajani S., Busse M., Yuan J., Lee G., Peale F.V., Klijn C., Bourgon R., Kaminker J.S., Neve R.M.
A resource for cell line authentication, annotation and quality control.
Nature 520:307-311(2015)
PubMed=25926053; DOI=10.1038/ncomms8002
Medico E., Russo M., Picco G., Cancelliere C., Valtorta E., Corti G., Buscarino M., Isella C., Lamba S., Martinoglio B., Veronese S., Siena S., Sartore-Bianchi A., Beccuti M., Mottolese M., Linnebacher M., Cordero F., Di Nicolantonio F., Bardelli A.
The molecular landscape of colorectal cancer cell lines unveils clinically actionable kinase targets.
Nat. Commun. 6:7002.1-7002.10(2015)
PubMed=25944804; DOI=10.1158/1078-0432.CCR-14-2457
Bazzocco S., Dopeso H., Carton-Garcia F., Macaya I., Andretta E., Chionh F., Rodrigues P., Garrido M., Alazzouzi H., Nieto R., Sanchez A., Schwartz S. Jr., Bilic J., Mariadason J.M., Arango D.
Highly expressed genes in rapidly proliferating tumor cells as new targets for colorectal cancer treatment.
Clin. Cancer Res. 21:3695-3704(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5; PMCID=PMC4653878
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=29787047; DOI=10.1007/978-3-319-16104-4_11
Martinez-Maqueda D., Miralles B., Recio I.
HT29 cell line.
(In book chapter) The impact of food bioactives on health. In vitro and ex vivo models; Verhoeckx K., Cotter P., Lopez-Exposito I., Kleiveland C., Lea T., Mackie A., Requena T., Swiatecka D., Wichers H. (eds.); pp.113-124; Springer; Cham; Switzerland (2015)
PubMed=26537799; DOI=10.1074/mcp.M115.051235; PMCID=PMC4762531
Holst S., Deuss A.J.M., van Pelt G.W., van Vliet S.J., Garcia-Vallejo J.J., Koeleman C.A.M., Deelder A.M., Mesker W.E., Tollenaar R.A.E.M., Rombouts Y., Wuhrer M.
N-glycosylation profiling of colorectal cancer cell lines reveals association of fucosylation with differentiation and caudal type homebox 1 (CDX1)/villin mRNA expression.
Mol. Cell. Proteomics 15:124-140(2016)
PubMed=27377824; DOI=10.1038/sdata.2016.52; PMCID=PMC4932877
Mestdagh P., Lefever S., Volders P.-J., Derveaux S., Hellemans J., Vandesompele J.
Long non-coding RNA expression profiling in the NCI60 cancer cell line panel using high-throughput RT-qPCR.
Sci. Data 3:160052-160052(2016)
PubMed=27397505; DOI=10.1016/j.cell.2016.06.017; PMCID=PMC4967469
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., Aben N., Goncalves E., Barthorpe S., Lightfoot H., Cokelaer T., Greninger P., van Dyk E., Chang H., de Silva H., Heyn H., Deng X.-M., Egan R.K., Liu Q.-S., Miroo T., Mitropoulos X., Richardson L., Wang J.-H., Zhang T.-H., Moran S., Sayols S., Soleimani M., Tamborero D., Lopez-Bigas N., Ross-Macdonald P., Esteller M., Gray N.S., Haber D.A., Stratton M.R., Benes C.H., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Garnett M.J.
A landscape of pharmacogenomic interactions in cancer.
Cell 166:740-754(2016)
PubMed=27807467; DOI=10.1186/s13100-016-0078-4; PMCID=PMC5087121
Zampella J.G., Rodic N., Yang W.R., Huang C.R.L., Welch J., Gnanakkan V.P., Cornish T.C., Boeke J.D., Burns K.H.
A map of mobile DNA insertions in the NCI-60 human cancer cell panel.
Mob. DNA 7:20.1-20.11(2016)
PubMed=28179481; DOI=10.1158/1535-7163.MCT-16-0578
Tanaka N., Mashima T., Mizutani A., Sato A., Aoyama A., Gong B., Yoshida H., Muramatsu Y., Nakata K., Matsuura M., Katayama R., Nagayama S., Fujita N., Sugimoto Y., Seimiya H.
APC mutations as a potential biomarker for sensitivity to tankyrase inhibitors in colorectal cancer.
Mol. Cancer Ther. 16:752-762(2017)
PubMed=28192450; DOI=10.1371/journal.pone.0171435; PMCID=PMC5305277
Fasterius E., Raso C., Kennedy S.A., Rauch N., Lundin P., Kolch W., Uhlen M., Al-Khalili Szigyarto C.
A novel RNA sequencing data analysis method for cell line authentication.
PLoS ONE 12:E0171435-E0171435(2017)
PubMed=28196595; DOI=10.1016/j.ccell.2017.01.005; PMCID=PMC5501076
Li J., Zhao W., Akbani R., Liu W.-B., Ju Z.-L., Ling S.-Y., Vellano C.P., Roebuck P., Yu Q.-H., Eterovic A.K., Byers L.A., Davies M.A., Deng W.-L., Gopal Y.N.V., Chen G., von Euw E.M., Slamon D.J., Conklin D., Heymach J.V., Gazdar A.F., Minna J.D., Myers J.N., Lu Y.-L., Mills G.B., Liang H.
Characterization of human cancer cell lines by reverse-phase protein arrays.
Cancer Cell 31:225-239(2017)
PubMed=28683746; DOI=10.1186/s12943-017-0691-y; PMCID=PMC5498998
Berg K.C.G., Eide P.W., Eilertsen I.A., Johannessen B., Bruun J., Danielsen S.A., Bjornslett M., Meza-Zepeda L.A., Eknaes M., Lind G.E., Myklebost O., Skotheim R.I., Sveen A., Lothe R.A.
Multi-omics of 34 colorectal cancer cell lines -- a resource for biomedical studies.
Mol. Cancer 16:116.1-116.16(2017)
PubMed=28854368; DOI=10.1016/j.celrep.2017.08.010; PMCID=PMC5583477
Roumeliotis T.I., Williams S.P., Goncalves E., Alsinet C., Del Castillo Velasco-Herrera M., Aben N., Ghavidel F.Z., Michaut M., Schubert M., Price S., Wright J.C., Yu L., Yang M., Dienstmann R., Guinney J.H., Beltrao P., Brazma A., Pardo M., Stegle O., Adams D.J., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Choudhary J.S.
Genomic determinants of protein abundance variation in colorectal cancer cells.
Cell Rep. 20:2201-2214(2017)
PubMed=29101300; DOI=10.15252/msb.20177701; PMCID=PMC5731344
Frejno M., Zenezini Chiozzi R., Wilhelm M., Koch H., Zheng R.-S., Klaeger S., Ruprecht B., Meng C., Kramer K., Jarzab A., Heinzlmeir S., Johnstone E., Domingo E., Kerr D.J., Jesinghaus M., Slotta-Huspenina J., Weichert W., Knapp S., Feller S.M., Kuster B.
Pharmacoproteomic characterisation of human colon and rectal cancer.
Mol. Syst. Biol. 13:951-951(2017)
PubMed=29207035; DOI=10.3892/ijo.2017.4206
Olejniczak A., Szarynska M., Kmiec Z.
In vitro characterization of spheres derived from colorectal cancer cell lines.
Int. J. Oncol. 52:599-612(2018)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747; PMCID=PMC6445675
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=30971826; DOI=10.1038/s41586-019-1103-9
Behan F.M., Iorio F., Picco G., Goncalves E., Beaver C.M., Migliardi G., Santos R., Rao Y., Sassi F., Pinnelli M., Ansari R., Harper S., Jackson D.A., McRae R., Pooley R., Wilkinson P., van der Meer D.J., Dow D., Buser-Doepner C.A., Bertotti A., Trusolino L., Stronach E.A., Saez-Rodriguez J., Yusa K., Garnett M.J.
Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens.
Nature 568:511-516(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3; PMCID=PMC6697103
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. 3rd, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=31978347; DOI=10.1016/j.cell.2019.12.023; PMCID=PMC7339254
Nusinow D.P., Szpyt J., Ghandi M., Rose C.M., McDonald E.R. 3rd, Kalocsay M., Jane-Valbuena J., Gelfand E.T., Schweppe D.K., Jedrychowski M.P., Golji J., Porter D.A., Rejtar T., Wang Y.K., Kryukov G.V., Stegmeier F., Erickson B.K., Garraway L.A., Sellers W.R., Gygi S.P.
Quantitative proteomics of the Cancer Cell Line Encyclopedia.
Cell 180:387-402.e16(2020)
PubMed=32172478; DOI=10.1007/s12253-020-00805-3
Xu Y.-T., Zhang L., Wang Q.-L., Zheng M.-J.
Comparison of different colorectal cancer with liver metastases models using six colorectal cancer cell lines.
Pathol. Oncol. R"